How to Write Limitations of the Study (with examples)
This blog emphasizes the importance of recognizing and effectively writing about limitations in research. It discusses the types of limitations, their significance, and provides guidelines for writing about them, highlighting their role in advancing scholarly research.
Updated on August 24, 2023

No matter how well thought out, every research endeavor encounters challenges. There is simply no way to predict all possible variances throughout the process.
These uncharted boundaries and abrupt constraints are known as limitations in research . Identifying and acknowledging limitations is crucial for conducting rigorous studies. Limitations provide context and shed light on gaps in the prevailing inquiry and literature.
This article explores the importance of recognizing limitations and discusses how to write them effectively. By interpreting limitations in research and considering prevalent examples, we aim to reframe the perception from shameful mistakes to respectable revelations.

What are limitations in research?
In the clearest terms, research limitations are the practical or theoretical shortcomings of a study that are often outside of the researcher’s control . While these weaknesses limit the generalizability of a study’s conclusions, they also present a foundation for future research.
Sometimes limitations arise from tangible circumstances like time and funding constraints, or equipment and participant availability. Other times the rationale is more obscure and buried within the research design. Common types of limitations and their ramifications include:
- Theoretical: limits the scope, depth, or applicability of a study.
- Methodological: limits the quality, quantity, or diversity of the data.
- Empirical: limits the representativeness, validity, or reliability of the data.
- Analytical: limits the accuracy, completeness, or significance of the findings.
- Ethical: limits the access, consent, or confidentiality of the data.
Regardless of how, when, or why they arise, limitations are a natural part of the research process and should never be ignored . Like all other aspects, they are vital in their own purpose.
Why is identifying limitations important?
Whether to seek acceptance or avoid struggle, humans often instinctively hide flaws and mistakes. Merging this thought process into research by attempting to hide limitations, however, is a bad idea. It has the potential to negate the validity of outcomes and damage the reputation of scholars.
By identifying and addressing limitations throughout a project, researchers strengthen their arguments and curtail the chance of peer censure based on overlooked mistakes. Pointing out these flaws shows an understanding of variable limits and a scrupulous research process.
Showing awareness of and taking responsibility for a project’s boundaries and challenges validates the integrity and transparency of a researcher. It further demonstrates the researchers understand the applicable literature and have thoroughly evaluated their chosen research methods.
Presenting limitations also benefits the readers by providing context for research findings. It guides them to interpret the project’s conclusions only within the scope of very specific conditions. By allowing for an appropriate generalization of the findings that is accurately confined by research boundaries and is not too broad, limitations boost a study’s credibility .
Limitations are true assets to the research process. They highlight opportunities for future research. When researchers identify the limitations of their particular approach to a study question, they enable precise transferability and improve chances for reproducibility.
Simply stating a project’s limitations is not adequate for spurring further research, though. To spark the interest of other researchers, these acknowledgements must come with thorough explanations regarding how the limitations affected the current study and how they can potentially be overcome with amended methods.
How to write limitations
Typically, the information about a study’s limitations is situated either at the beginning of the discussion section to provide context for readers or at the conclusion of the discussion section to acknowledge the need for further research. However, it varies depending upon the target journal or publication guidelines.
Don’t hide your limitations
It is also important to not bury a limitation in the body of the paper unless it has a unique connection to a topic in that section. If so, it needs to be reiterated with the other limitations or at the conclusion of the discussion section. Wherever it is included in the manuscript, ensure that the limitations section is prominently positioned and clearly introduced.
While maintaining transparency by disclosing limitations means taking a comprehensive approach, it is not necessary to discuss everything that could have potentially gone wrong during the research study. If there is no commitment to investigation in the introduction, it is unnecessary to consider the issue a limitation to the research. Wholly consider the term ‘limitations’ and ask, “Did it significantly change or limit the possible outcomes?” Then, qualify the occurrence as either a limitation to include in the current manuscript or as an idea to note for other projects.
Writing limitations
Once the limitations are concretely identified and it is decided where they will be included in the paper, researchers are ready for the writing task. Including only what is pertinent, keeping explanations detailed but concise, and employing the following guidelines is key for crafting valuable limitations:
1) Identify and describe the limitations : Clearly introduce the limitation by classifying its form and specifying its origin. For example:
- An unintentional bias encountered during data collection
- An intentional use of unplanned post-hoc data analysis
2) Explain the implications : Describe how the limitation potentially influences the study’s findings and how the validity and generalizability are subsequently impacted. Provide examples and evidence to support claims of the limitations’ effects without making excuses or exaggerating their impact. Overall, be transparent and objective in presenting the limitations, without undermining the significance of the research.
3) Provide alternative approaches for future studies : Offer specific suggestions for potential improvements or avenues for further investigation. Demonstrate a proactive approach by encouraging future research that addresses the identified gaps and, therefore, expands the knowledge base.
Whether presenting limitations as an individual section within the manuscript or as a subtopic in the discussion area, authors should use clear headings and straightforward language to facilitate readability. There is no need to complicate limitations with jargon, computations, or complex datasets.
Examples of common limitations
Limitations are generally grouped into two categories , methodology and research process .
Methodology limitations
Methodology may include limitations due to:
- Sample size
- Lack of available or reliable data
- Lack of prior research studies on the topic
- Measure used to collect the data
- Self-reported data

The researcher is addressing how the large sample size requires a reassessment of the measures used to collect and analyze the data.
Research process limitations
Limitations during the research process may arise from:
- Access to information
- Longitudinal effects
- Cultural and other biases
- Language fluency
- Time constraints

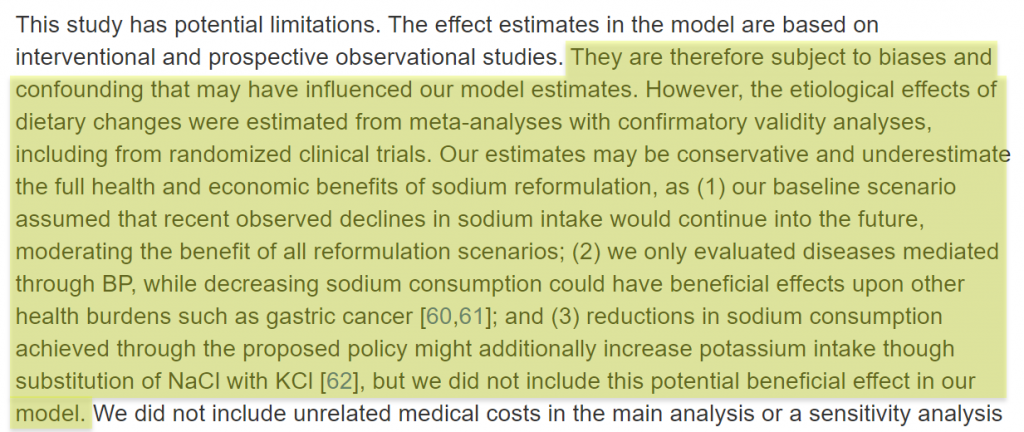
The author is pointing out that the model’s estimates are based on potentially biased observational studies.
Final thoughts
Successfully proving theories and touting great achievements are only two very narrow goals of scholarly research. The true passion and greatest efforts of researchers comes more in the form of confronting assumptions and exploring the obscure.
In many ways, recognizing and sharing the limitations of a research study both allows for and encourages this type of discovery that continuously pushes research forward. By using limitations to provide a transparent account of the project's boundaries and to contextualize the findings, researchers pave the way for even more robust and impactful research in the future.
Charla Viera, MS
See our "Privacy Policy"
Ensure your structure and ideas are consistent and clearly communicated
Pair your Premium Editing with our add-on service Presubmission Review for an overall assessment of your manuscript.
21 Research Limitations Examples

Chris Drew (PhD)
Dr. Chris Drew is the founder of the Helpful Professor. He holds a PhD in education and has published over 20 articles in scholarly journals. He is the former editor of the Journal of Learning Development in Higher Education. [Image Descriptor: Photo of Chris]
Learn about our Editorial Process
Research limitations refer to the potential weaknesses inherent in a study. All studies have limitations of some sort, meaning declaring limitations doesn’t necessarily need to be a bad thing, so long as your declaration of limitations is well thought-out and explained.
Rarely is a study perfect. Researchers have to make trade-offs when developing their studies, which are often based upon practical considerations such as time and monetary constraints, weighing the breadth of participants against the depth of insight, and choosing one methodology or another.
In research, studies can have limitations such as limited scope, researcher subjectivity, and lack of available research tools.
Acknowledging the limitations of your study should be seen as a strength. It demonstrates your willingness for transparency, humility, and submission to the scientific method and can bolster the integrity of the study. It can also inform future research direction.
Typically, scholars will explore the limitations of their study in either their methodology section, their conclusion section, or both.
Research Limitations Examples
Qualitative and quantitative research offer different perspectives and methods in exploring phenomena, each with its own strengths and limitations. So, I’ve split the limitations examples sections into qualitative and quantitative below.
Qualitative Research Limitations
Qualitative research seeks to understand phenomena in-depth and in context. It focuses on the ‘why’ and ‘how’ questions.
It’s often used to explore new or complex issues, and it provides rich, detailed insights into participants’ experiences, behaviors, and attitudes. However, these strengths also create certain limitations, as explained below.
1. Subjectivity
Qualitative research often requires the researcher to interpret subjective data. One researcher may examine a text and identify different themes or concepts as more dominant than others.
Close qualitative readings of texts are necessarily subjective – and while this may be a limitation, qualitative researchers argue this is the best way to deeply understand everything in context.
Suggested Solution and Response: To minimize subjectivity bias, you could consider cross-checking your own readings of themes and data against other scholars’ readings and interpretations. This may involve giving the raw data to a supervisor or colleague and asking them to code the data separately, then coming together to compare and contrast results.
2. Researcher Bias
The concept of researcher bias is related to, but slightly different from, subjectivity.
Researcher bias refers to the perspectives and opinions you bring with you when doing your research.
For example, a researcher who is explicitly of a certain philosophical or political persuasion may bring that persuasion to bear when interpreting data.
In many scholarly traditions, we will attempt to minimize researcher bias through the utilization of clear procedures that are set out in advance or through the use of statistical analysis tools.
However, in other traditions, such as in postmodern feminist research , declaration of bias is expected, and acknowledgment of bias is seen as a positive because, in those traditions, it is believed that bias cannot be eliminated from research, so instead, it is a matter of integrity to present it upfront.
Suggested Solution and Response: Acknowledge the potential for researcher bias and, depending on your theoretical framework , accept this, or identify procedures you have taken to seek a closer approximation to objectivity in your coding and analysis.
3. Generalizability
If you’re struggling to find a limitation to discuss in your own qualitative research study, then this one is for you: all qualitative research, of all persuasions and perspectives, cannot be generalized.
This is a core feature that sets qualitative data and quantitative data apart.
The point of qualitative data is to select case studies and similarly small corpora and dig deep through in-depth analysis and thick description of data.
Often, this will also mean that you have a non-randomized sample size.
While this is a positive – you’re going to get some really deep, contextualized, interesting insights – it also means that the findings may not be generalizable to a larger population that may not be representative of the small group of people in your study.
Suggested Solution and Response: Suggest future studies that take a quantitative approach to the question.
4. The Hawthorne Effect
The Hawthorne effect refers to the phenomenon where research participants change their ‘observed behavior’ when they’re aware that they are being observed.
This effect was first identified by Elton Mayo who conducted studies of the effects of various factors ton workers’ productivity. He noticed that no matter what he did – turning up the lights, turning down the lights, etc. – there was an increase in worker outputs compared to prior to the study taking place.
Mayo realized that the mere act of observing the workers made them work harder – his observation was what was changing behavior.
So, if you’re looking for a potential limitation to name for your observational research study , highlight the possible impact of the Hawthorne effect (and how you could reduce your footprint or visibility in order to decrease its likelihood).
Suggested Solution and Response: Highlight ways you have attempted to reduce your footprint while in the field, and guarantee anonymity to your research participants.
5. Replicability
Quantitative research has a great benefit in that the studies are replicable – a researcher can get a similar sample size, duplicate the variables, and re-test a study. But you can’t do that in qualitative research.
Qualitative research relies heavily on context – a specific case study or specific variables that make a certain instance worthy of analysis. As a result, it’s often difficult to re-enter the same setting with the same variables and repeat the study.
Furthermore, the individual researcher’s interpretation is more influential in qualitative research, meaning even if a new researcher enters an environment and makes observations, their observations may be different because subjectivity comes into play much more. This doesn’t make the research bad necessarily (great insights can be made in qualitative research), but it certainly does demonstrate a weakness of qualitative research.
6. Limited Scope
“Limited scope” is perhaps one of the most common limitations listed by researchers – and while this is often a catch-all way of saying, “well, I’m not studying that in this study”, it’s also a valid point.
No study can explore everything related to a topic. At some point, we have to make decisions about what’s included in the study and what is excluded from the study.
So, you could say that a limitation of your study is that it doesn’t look at an extra variable or concept that’s certainly worthy of study but will have to be explored in your next project because this project has a clearly and narrowly defined goal.
Suggested Solution and Response: Be clear about what’s in and out of the study when writing your research question.
7. Time Constraints
This is also a catch-all claim you can make about your research project: that you would have included more people in the study, looked at more variables, and so on. But you’ve got to submit this thing by the end of next semester! You’ve got time constraints.
And time constraints are a recognized reality in all research.
But this means you’ll need to explain how time has limited your decisions. As with “limited scope”, this may mean that you had to study a smaller group of subjects, limit the amount of time you spent in the field, and so forth.
Suggested Solution and Response: Suggest future studies that will build on your current work, possibly as a PhD project.
8. Resource Intensiveness
Qualitative research can be expensive due to the cost of transcription, the involvement of trained researchers, and potential travel for interviews or observations.
So, resource intensiveness is similar to the time constraints concept. If you don’t have the funds, you have to make decisions about which tools to use, which statistical software to employ, and how many research assistants you can dedicate to the study.
Suggested Solution and Response: Suggest future studies that will gain more funding on the back of this ‘ exploratory study ‘.
9. Coding Difficulties
Data analysis in qualitative research often involves coding, which can be subjective and complex, especially when dealing with ambiguous or contradicting data.
After naming this as a limitation in your research, it’s important to explain how you’ve attempted to address this. Some ways to ‘limit the limitation’ include:
- Triangulation: Have 2 other researchers code the data as well and cross-check your results with theirs to identify outliers that may need to be re-examined, debated with the other researchers, or removed altogether.
- Procedure: Use a clear coding procedure to demonstrate reliability in your coding process. I personally use the thematic network analysis method outlined in this academic article by Attride-Stirling (2001).
Suggested Solution and Response: Triangulate your coding findings with colleagues, and follow a thematic network analysis procedure.
10. Risk of Non-Responsiveness
There is always a risk in research that research participants will be unwilling or uncomfortable sharing their genuine thoughts and feelings in the study.
This is particularly true when you’re conducting research on sensitive topics, politicized topics, or topics where the participant is expressing vulnerability .
This is similar to the Hawthorne effect (aka participant bias), where participants change their behaviors in your presence; but it goes a step further, where participants actively hide their true thoughts and feelings from you.
Suggested Solution and Response: One way to manage this is to try to include a wider group of people with the expectation that there will be non-responsiveness from some participants.
11. Risk of Attrition
Attrition refers to the process of losing research participants throughout the study.
This occurs most commonly in longitudinal studies , where a researcher must return to conduct their analysis over spaced periods of time, often over a period of years.
Things happen to people over time – they move overseas, their life experiences change, they get sick, change their minds, and even die. The more time that passes, the greater the risk of attrition.
Suggested Solution and Response: One way to manage this is to try to include a wider group of people with the expectation that there will be attrition over time.
12. Difficulty in Maintaining Confidentiality and Anonymity
Given the detailed nature of qualitative data , ensuring participant anonymity can be challenging.
If you have a sensitive topic in a specific case study, even anonymizing research participants sometimes isn’t enough. People might be able to induce who you’re talking about.
Sometimes, this will mean you have to exclude some interesting data that you collected from your final report. Confidentiality and anonymity come before your findings in research ethics – and this is a necessary limiting factor.
Suggested Solution and Response: Highlight the efforts you have taken to anonymize data, and accept that confidentiality and accountability place extremely important constraints on academic research.
13. Difficulty in Finding Research Participants
A study that looks at a very specific phenomenon or even a specific set of cases within a phenomenon means that the pool of potential research participants can be very low.
Compile on top of this the fact that many people you approach may choose not to participate, and you could end up with a very small corpus of subjects to explore. This may limit your ability to make complete findings, even in a quantitative sense.
You may need to therefore limit your research question and objectives to something more realistic.
Suggested Solution and Response: Highlight that this is going to limit the study’s generalizability significantly.
14. Ethical Limitations
Ethical limitations refer to the things you cannot do based on ethical concerns identified either by yourself or your institution’s ethics review board.
This might include threats to the physical or psychological well-being of your research subjects, the potential of releasing data that could harm a person’s reputation, and so on.
Furthermore, even if your study follows all expected standards of ethics, you still, as an ethical researcher, need to allow a research participant to pull out at any point in time, after which you cannot use their data, which demonstrates an overlap between ethical constraints and participant attrition.
Suggested Solution and Response: Highlight that these ethical limitations are inevitable but important to sustain the integrity of the research.
For more on Qualitative Research, Explore my Qualitative Research Guide
Quantitative Research Limitations
Quantitative research focuses on quantifiable data and statistical, mathematical, or computational techniques. It’s often used to test hypotheses, assess relationships and causality, and generalize findings across larger populations.
Quantitative research is widely respected for its ability to provide reliable, measurable, and generalizable data (if done well!). Its structured methodology has strengths over qualitative research, such as the fact it allows for replication of the study, which underpins the validity of the research.
However, this approach is not without it limitations, explained below.
1. Over-Simplification
Quantitative research is powerful because it allows you to measure and analyze data in a systematic and standardized way. However, one of its limitations is that it can sometimes simplify complex phenomena or situations.
In other words, it might miss the subtleties or nuances of the research subject.
For example, if you’re studying why people choose a particular diet, a quantitative study might identify factors like age, income, or health status. But it might miss other aspects, such as cultural influences or personal beliefs, that can also significantly impact dietary choices.
When writing about this limitation, you can say that your quantitative approach, while providing precise measurements and comparisons, may not capture the full complexity of your subjects of study.
Suggested Solution and Response: Suggest a follow-up case study using the same research participants in order to gain additional context and depth.
2. Lack of Context
Another potential issue with quantitative research is that it often focuses on numbers and statistics at the expense of context or qualitative information.
Let’s say you’re studying the effect of classroom size on student performance. You might find that students in smaller classes generally perform better. However, this doesn’t take into account other variables, like teaching style , student motivation, or family support.
When describing this limitation, you might say, “Although our research provides important insights into the relationship between class size and student performance, it does not incorporate the impact of other potentially influential variables. Future research could benefit from a mixed-methods approach that combines quantitative analysis with qualitative insights.”
3. Applicability to Real-World Settings
Oftentimes, experimental research takes place in controlled environments to limit the influence of outside factors.
This control is great for isolation and understanding the specific phenomenon but can limit the applicability or “external validity” of the research to real-world settings.
For example, if you conduct a lab experiment to see how sleep deprivation impacts cognitive performance, the sterile, controlled lab environment might not reflect real-world conditions where people are dealing with multiple stressors.
Therefore, when explaining the limitations of your quantitative study in your methodology section, you could state:
“While our findings provide valuable information about [topic], the controlled conditions of the experiment may not accurately represent real-world scenarios where extraneous variables will exist. As such, the direct applicability of our results to broader contexts may be limited.”
Suggested Solution and Response: Suggest future studies that will engage in real-world observational research, such as ethnographic research.
4. Limited Flexibility
Once a quantitative study is underway, it can be challenging to make changes to it. This is because, unlike in grounded research, you’re putting in place your study in advance, and you can’t make changes part-way through.
Your study design, data collection methods, and analysis techniques need to be decided upon before you start collecting data.
For example, if you are conducting a survey on the impact of social media on teenage mental health, and halfway through, you realize that you should have included a question about their screen time, it’s generally too late to add it.
When discussing this limitation, you could write something like, “The structured nature of our quantitative approach allows for consistent data collection and analysis but also limits our flexibility to adapt and modify the research process in response to emerging insights and ideas.”
Suggested Solution and Response: Suggest future studies that will use mixed-methods or qualitative research methods to gain additional depth of insight.
5. Risk of Survey Error
Surveys are a common tool in quantitative research, but they carry risks of error.
There can be measurement errors (if a question is misunderstood), coverage errors (if some groups aren’t adequately represented), non-response errors (if certain people don’t respond), and sampling errors (if your sample isn’t representative of the population).
For instance, if you’re surveying college students about their study habits , but only daytime students respond because you conduct the survey during the day, your results will be skewed.
In discussing this limitation, you might say, “Despite our best efforts to develop a comprehensive survey, there remains a risk of survey error, including measurement, coverage, non-response, and sampling errors. These could potentially impact the reliability and generalizability of our findings.”
Suggested Solution and Response: Suggest future studies that will use other survey tools to compare and contrast results.
6. Limited Ability to Probe Answers
With quantitative research, you typically can’t ask follow-up questions or delve deeper into participants’ responses like you could in a qualitative interview.
For instance, imagine you are surveying 500 students about study habits in a questionnaire. A respondent might indicate that they study for two hours each night. You might want to follow up by asking them to elaborate on what those study sessions involve or how effective they feel their habits are.
However, quantitative research generally disallows this in the way a qualitative semi-structured interview could.
When discussing this limitation, you might write, “Given the structured nature of our survey, our ability to probe deeper into individual responses is limited. This means we may not fully understand the context or reasoning behind the responses, potentially limiting the depth of our findings.”
Suggested Solution and Response: Suggest future studies that engage in mixed-method or qualitative methodologies to address the issue from another angle.
7. Reliance on Instruments for Data Collection
In quantitative research, the collection of data heavily relies on instruments like questionnaires, surveys, or machines.
The limitation here is that the data you get is only as good as the instrument you’re using. If the instrument isn’t designed or calibrated well, your data can be flawed.
For instance, if you’re using a questionnaire to study customer satisfaction and the questions are vague, confusing, or biased, the responses may not accurately reflect the customers’ true feelings.
When discussing this limitation, you could say, “Our study depends on the use of questionnaires for data collection. Although we have put significant effort into designing and testing the instrument, it’s possible that inaccuracies or misunderstandings could potentially affect the validity of the data collected.”
Suggested Solution and Response: Suggest future studies that will use different instruments but examine the same variables to triangulate results.
8. Time and Resource Constraints (Specific to Quantitative Research)
Quantitative research can be time-consuming and resource-intensive, especially when dealing with large samples.
It often involves systematic sampling, rigorous design, and sometimes complex statistical analysis.
If resources and time are limited, it can restrict the scale of your research, the techniques you can employ, or the extent of your data analysis.
For example, you may want to conduct a nationwide survey on public opinion about a certain policy. However, due to limited resources, you might only be able to survey people in one city.
When writing about this limitation, you could say, “Given the scope of our research and the resources available, we are limited to conducting our survey within one city, which may not fully represent the nationwide public opinion. Hence, the generalizability of the results may be limited.”
Suggested Solution and Response: Suggest future studies that will have more funding or longer timeframes.
How to Discuss Your Research Limitations
1. in your research proposal and methodology section.
In the research proposal, which will become the methodology section of your dissertation, I would recommend taking the four following steps, in order:
- Be Explicit about your Scope – If you limit the scope of your study in your research question, aims, and objectives, then you can set yourself up well later in the methodology to say that certain questions are “outside the scope of the study.” For example, you may identify the fact that the study doesn’t address a certain variable, but you can follow up by stating that the research question is specifically focused on the variable that you are examining, so this limitation would need to be looked at in future studies.
- Acknowledge the Limitation – Acknowledging the limitations of your study demonstrates reflexivity and humility and can make your research more reliable and valid. It also pre-empts questions the people grading your paper may have, so instead of them down-grading you for your limitations; they will congratulate you on explaining the limitations and how you have addressed them!
- Explain your Decisions – You may have chosen your approach (despite its limitations) for a very specific reason. This might be because your approach remains, on balance, the best one to answer your research question. Or, it might be because of time and monetary constraints that are outside of your control.
- Highlight the Strengths of your Approach – Conclude your limitations section by strongly demonstrating that, despite limitations, you’ve worked hard to minimize the effects of the limitations and that you have chosen your specific approach and methodology because it’s also got some terrific strengths. Name the strengths.
Overall, you’ll want to acknowledge your own limitations but also explain that the limitations don’t detract from the value of your study as it stands.
2. In the Conclusion Section or Chapter
In the conclusion of your study, it is generally expected that you return to a discussion of the study’s limitations. Here, I recommend the following steps:
- Acknowledge issues faced – After completing your study, you will be increasingly aware of issues you may have faced that, if you re-did the study, you may have addressed earlier in order to avoid those issues. Acknowledge these issues as limitations, and frame them as recommendations for subsequent studies.
- Suggest further research – Scholarly research aims to fill gaps in the current literature and knowledge. Having established your expertise through your study, suggest lines of inquiry for future researchers. You could state that your study had certain limitations, and “future studies” can address those limitations.
- Suggest a mixed methods approach – Qualitative and quantitative research each have pros and cons. So, note those ‘cons’ of your approach, then say the next study should approach the topic using the opposite methodology or could approach it using a mixed-methods approach that could achieve the benefits of quantitative studies with the nuanced insights of associated qualitative insights as part of an in-study case-study.
Overall, be clear about both your limitations and how those limitations can inform future studies.
In sum, each type of research method has its own strengths and limitations. Qualitative research excels in exploring depth, context, and complexity, while quantitative research excels in examining breadth, generalizability, and quantifiable measures. Despite their individual limitations, each method contributes unique and valuable insights, and researchers often use them together to provide a more comprehensive understanding of the phenomenon being studied.
Attride-Stirling, J. (2001). Thematic networks: an analytic tool for qualitative research. Qualitative research , 1 (3), 385-405. ( Source )
Atkinson, P., Delamont, S., Cernat, A., Sakshaug, J., & Williams, R. A. (2021). SAGE research methods foundations . London: Sage Publications.
Clark, T., Foster, L., Bryman, A., & Sloan, L. (2021). Bryman’s social research methods . Oxford: Oxford University Press.
Köhler, T., Smith, A., & Bhakoo, V. (2022). Templates in qualitative research methods: Origins, limitations, and new directions. Organizational Research Methods , 25 (2), 183-210. ( Source )
Lenger, A. (2019). The rejection of qualitative research methods in economics. Journal of Economic Issues , 53 (4), 946-965. ( Source )
Taherdoost, H. (2022). What are different research approaches? Comprehensive review of qualitative, quantitative, and mixed method research, their applications, types, and limitations. Journal of Management Science & Engineering Research , 5 (1), 53-63. ( Source )
Walliman, N. (2021). Research methods: The basics . New York: Routledge.

- Chris Drew (PhD) https://helpfulprofessor.com/author/chris-drew-phd/ 19 Top Cognitive Psychology Theories (Explained)
- Chris Drew (PhD) https://helpfulprofessor.com/author/chris-drew-phd/ 119 Bloom’s Taxonomy Examples
- Chris Drew (PhD) https://helpfulprofessor.com/author/chris-drew-phd/ All 6 Levels of Understanding (on Bloom’s Taxonomy)
- Chris Drew (PhD) https://helpfulprofessor.com/author/chris-drew-phd/ 15 Self-Actualization Examples (Maslow's Hierarchy)
Leave a Comment Cancel Reply
Your email address will not be published. Required fields are marked *
- Privacy Policy

Home » Limitations in Research – Types, Examples and Writing Guide
Limitations in Research – Types, Examples and Writing Guide
Table of Contents

Limitations in Research
Limitations in research refer to the factors that may affect the results, conclusions , and generalizability of a study. These limitations can arise from various sources, such as the design of the study, the sampling methods used, the measurement tools employed, and the limitations of the data analysis techniques.
Types of Limitations in Research
Types of Limitations in Research are as follows:
Sample Size Limitations
This refers to the size of the group of people or subjects that are being studied. If the sample size is too small, then the results may not be representative of the population being studied. This can lead to a lack of generalizability of the results.
Time Limitations
Time limitations can be a constraint on the research process . This could mean that the study is unable to be conducted for a long enough period of time to observe the long-term effects of an intervention, or to collect enough data to draw accurate conclusions.
Selection Bias
This refers to a type of bias that can occur when the selection of participants in a study is not random. This can lead to a biased sample that is not representative of the population being studied.
Confounding Variables
Confounding variables are factors that can influence the outcome of a study, but are not being measured or controlled for. These can lead to inaccurate conclusions or a lack of clarity in the results.
Measurement Error
This refers to inaccuracies in the measurement of variables, such as using a faulty instrument or scale. This can lead to inaccurate results or a lack of validity in the study.
Ethical Limitations
Ethical limitations refer to the ethical constraints placed on research studies. For example, certain studies may not be allowed to be conducted due to ethical concerns, such as studies that involve harm to participants.
Examples of Limitations in Research
Some Examples of Limitations in Research are as follows:
Research Title: “The Effectiveness of Machine Learning Algorithms in Predicting Customer Behavior”
Limitations:
- The study only considered a limited number of machine learning algorithms and did not explore the effectiveness of other algorithms.
- The study used a specific dataset, which may not be representative of all customer behaviors or demographics.
- The study did not consider the potential ethical implications of using machine learning algorithms in predicting customer behavior.
Research Title: “The Impact of Online Learning on Student Performance in Computer Science Courses”
- The study was conducted during the COVID-19 pandemic, which may have affected the results due to the unique circumstances of remote learning.
- The study only included students from a single university, which may limit the generalizability of the findings to other institutions.
- The study did not consider the impact of individual differences, such as prior knowledge or motivation, on student performance in online learning environments.
Research Title: “The Effect of Gamification on User Engagement in Mobile Health Applications”
- The study only tested a specific gamification strategy and did not explore the effectiveness of other gamification techniques.
- The study relied on self-reported measures of user engagement, which may be subject to social desirability bias or measurement errors.
- The study only included a specific demographic group (e.g., young adults) and may not be generalizable to other populations with different preferences or needs.
How to Write Limitations in Research
When writing about the limitations of a research study, it is important to be honest and clear about the potential weaknesses of your work. Here are some tips for writing about limitations in research:
- Identify the limitations: Start by identifying the potential limitations of your research. These may include sample size, selection bias, measurement error, or other issues that could affect the validity and reliability of your findings.
- Be honest and objective: When describing the limitations of your research, be honest and objective. Do not try to minimize or downplay the limitations, but also do not exaggerate them. Be clear and concise in your description of the limitations.
- Provide context: It is important to provide context for the limitations of your research. For example, if your sample size was small, explain why this was the case and how it may have affected your results. Providing context can help readers understand the limitations in a broader context.
- Discuss implications : Discuss the implications of the limitations for your research findings. For example, if there was a selection bias in your sample, explain how this may have affected the generalizability of your findings. This can help readers understand the limitations in terms of their impact on the overall validity of your research.
- Provide suggestions for future research : Finally, provide suggestions for future research that can address the limitations of your study. This can help readers understand how your research fits into the broader field and can provide a roadmap for future studies.
Purpose of Limitations in Research
There are several purposes of limitations in research. Here are some of the most important ones:
- To acknowledge the boundaries of the study : Limitations help to define the scope of the research project and set realistic expectations for the findings. They can help to clarify what the study is not intended to address.
- To identify potential sources of bias: Limitations can help researchers identify potential sources of bias in their research design, data collection, or analysis. This can help to improve the validity and reliability of the findings.
- To provide opportunities for future research: Limitations can highlight areas for future research and suggest avenues for further exploration. This can help to advance knowledge in a particular field.
- To demonstrate transparency and accountability: By acknowledging the limitations of their research, researchers can demonstrate transparency and accountability to their readers, peers, and funders. This can help to build trust and credibility in the research community.
- To encourage critical thinking: Limitations can encourage readers to critically evaluate the study’s findings and consider alternative explanations or interpretations. This can help to promote a more nuanced and sophisticated understanding of the topic under investigation.
When to Write Limitations in Research
Limitations should be included in research when they help to provide a more complete understanding of the study’s results and implications. A limitation is any factor that could potentially impact the accuracy, reliability, or generalizability of the study’s findings.
It is important to identify and discuss limitations in research because doing so helps to ensure that the results are interpreted appropriately and that any conclusions drawn are supported by the available evidence. Limitations can also suggest areas for future research, highlight potential biases or confounding factors that may have affected the results, and provide context for the study’s findings.
Generally, limitations should be discussed in the conclusion section of a research paper or thesis, although they may also be mentioned in other sections, such as the introduction or methods. The specific limitations that are discussed will depend on the nature of the study, the research question being investigated, and the data that was collected.
Examples of limitations that might be discussed in research include sample size limitations, data collection methods, the validity and reliability of measures used, and potential biases or confounding factors that could have affected the results. It is important to note that limitations should not be used as a justification for poor research design or methodology, but rather as a way to enhance the understanding and interpretation of the study’s findings.
Importance of Limitations in Research
Here are some reasons why limitations are important in research:
- Enhances the credibility of research: Limitations highlight the potential weaknesses and threats to validity, which helps readers to understand the scope and boundaries of the study. This improves the credibility of research by acknowledging its limitations and providing a clear picture of what can and cannot be concluded from the study.
- Facilitates replication: By highlighting the limitations, researchers can provide detailed information about the study’s methodology, data collection, and analysis. This information helps other researchers to replicate the study and test the validity of the findings, which enhances the reliability of research.
- Guides future research : Limitations provide insights into areas for future research by identifying gaps or areas that require further investigation. This can help researchers to design more comprehensive and effective studies that build on existing knowledge.
- Provides a balanced view: Limitations help to provide a balanced view of the research by highlighting both strengths and weaknesses. This ensures that readers have a clear understanding of the study’s limitations and can make informed decisions about the generalizability and applicability of the findings.
Advantages of Limitations in Research
Here are some potential advantages of limitations in research:
- Focus : Limitations can help researchers focus their study on a specific area or population, which can make the research more relevant and useful.
- Realism : Limitations can make a study more realistic by reflecting the practical constraints and challenges of conducting research in the real world.
- Innovation : Limitations can spur researchers to be more innovative and creative in their research design and methodology, as they search for ways to work around the limitations.
- Rigor : Limitations can actually increase the rigor and credibility of a study, as researchers are forced to carefully consider the potential sources of bias and error, and address them to the best of their abilities.
- Generalizability : Limitations can actually improve the generalizability of a study by ensuring that it is not overly focused on a specific sample or situation, and that the results can be applied more broadly.
About the author
Muhammad Hassan
Researcher, Academic Writer, Web developer
You may also like

APA Table of Contents – Format and Example

Delimitations in Research – Types, Examples and...

Institutional Review Board – Application Sample...

Appendix in Research Paper – Examples and...

Research Paper – Structure, Examples and Writing...

Research Topics – Ideas and Examples
- USC Libraries
- Research Guides
Organizing Your Social Sciences Research Paper
- Limitations of the Study
- Purpose of Guide
- Design Flaws to Avoid
- Independent and Dependent Variables
- Glossary of Research Terms
- Reading Research Effectively
- Narrowing a Topic Idea
- Broadening a Topic Idea
- Extending the Timeliness of a Topic Idea
- Academic Writing Style
- Applying Critical Thinking
- Choosing a Title
- Making an Outline
- Paragraph Development
- Research Process Video Series
- Executive Summary
- The C.A.R.S. Model
- Background Information
- The Research Problem/Question
- Theoretical Framework
- Citation Tracking
- Content Alert Services
- Evaluating Sources
- Primary Sources
- Secondary Sources
- Tiertiary Sources
- Scholarly vs. Popular Publications
- Qualitative Methods
- Quantitative Methods
- Insiderness
- Using Non-Textual Elements
- Common Grammar Mistakes
- Writing Concisely
- Avoiding Plagiarism
- Footnotes or Endnotes?
- Further Readings
- Generative AI and Writing
- USC Libraries Tutorials and Other Guides
- Bibliography
The limitations of the study are those characteristics of design or methodology that impacted or influenced the interpretation of the findings from your research. Study limitations are the constraints placed on the ability to generalize from the results, to further describe applications to practice, and/or related to the utility of findings that are the result of the ways in which you initially chose to design the study or the method used to establish internal and external validity or the result of unanticipated challenges that emerged during the study.
Price, James H. and Judy Murnan. “Research Limitations and the Necessity of Reporting Them.” American Journal of Health Education 35 (2004): 66-67; Theofanidis, Dimitrios and Antigoni Fountouki. "Limitations and Delimitations in the Research Process." Perioperative Nursing 7 (September-December 2018): 155-163. .
Importance of...
Always acknowledge a study's limitations. It is far better that you identify and acknowledge your study’s limitations than to have them pointed out by your professor and have your grade lowered because you appeared to have ignored them or didn't realize they existed.
Keep in mind that acknowledgment of a study's limitations is an opportunity to make suggestions for further research. If you do connect your study's limitations to suggestions for further research, be sure to explain the ways in which these unanswered questions may become more focused because of your study.
Acknowledgment of a study's limitations also provides you with opportunities to demonstrate that you have thought critically about the research problem, understood the relevant literature published about it, and correctly assessed the methods chosen for studying the problem. A key objective of the research process is not only discovering new knowledge but also to confront assumptions and explore what we don't know.
Claiming limitations is a subjective process because you must evaluate the impact of those limitations . Don't just list key weaknesses and the magnitude of a study's limitations. To do so diminishes the validity of your research because it leaves the reader wondering whether, or in what ways, limitation(s) in your study may have impacted the results and conclusions. Limitations require a critical, overall appraisal and interpretation of their impact. You should answer the question: do these problems with errors, methods, validity, etc. eventually matter and, if so, to what extent?
Price, James H. and Judy Murnan. “Research Limitations and the Necessity of Reporting Them.” American Journal of Health Education 35 (2004): 66-67; Structure: How to Structure the Research Limitations Section of Your Dissertation. Dissertations and Theses: An Online Textbook. Laerd.com.
Descriptions of Possible Limitations
All studies have limitations . However, it is important that you restrict your discussion to limitations related to the research problem under investigation. For example, if a meta-analysis of existing literature is not a stated purpose of your research, it should not be discussed as a limitation. Do not apologize for not addressing issues that you did not promise to investigate in the introduction of your paper.
Here are examples of limitations related to methodology and the research process you may need to describe and discuss how they possibly impacted your results. Note that descriptions of limitations should be stated in the past tense because they were discovered after you completed your research.
Possible Methodological Limitations
- Sample size -- the number of the units of analysis you use in your study is dictated by the type of research problem you are investigating. Note that, if your sample size is too small, it will be difficult to find significant relationships from the data, as statistical tests normally require a larger sample size to ensure a representative distribution of the population and to be considered representative of groups of people to whom results will be generalized or transferred. Note that sample size is generally less relevant in qualitative research if explained in the context of the research problem.
- Lack of available and/or reliable data -- a lack of data or of reliable data will likely require you to limit the scope of your analysis, the size of your sample, or it can be a significant obstacle in finding a trend and a meaningful relationship. You need to not only describe these limitations but provide cogent reasons why you believe data is missing or is unreliable. However, don’t just throw up your hands in frustration; use this as an opportunity to describe a need for future research based on designing a different method for gathering data.
- Lack of prior research studies on the topic -- citing prior research studies forms the basis of your literature review and helps lay a foundation for understanding the research problem you are investigating. Depending on the currency or scope of your research topic, there may be little, if any, prior research on your topic. Before assuming this to be true, though, consult with a librarian! In cases when a librarian has confirmed that there is little or no prior research, you may be required to develop an entirely new research typology [for example, using an exploratory rather than an explanatory research design ]. Note again that discovering a limitation can serve as an important opportunity to identify new gaps in the literature and to describe the need for further research.
- Measure used to collect the data -- sometimes it is the case that, after completing your interpretation of the findings, you discover that the way in which you gathered data inhibited your ability to conduct a thorough analysis of the results. For example, you regret not including a specific question in a survey that, in retrospect, could have helped address a particular issue that emerged later in the study. Acknowledge the deficiency by stating a need for future researchers to revise the specific method for gathering data.
- Self-reported data -- whether you are relying on pre-existing data or you are conducting a qualitative research study and gathering the data yourself, self-reported data is limited by the fact that it rarely can be independently verified. In other words, you have to the accuracy of what people say, whether in interviews, focus groups, or on questionnaires, at face value. However, self-reported data can contain several potential sources of bias that you should be alert to and note as limitations. These biases become apparent if they are incongruent with data from other sources. These are: (1) selective memory [remembering or not remembering experiences or events that occurred at some point in the past]; (2) telescoping [recalling events that occurred at one time as if they occurred at another time]; (3) attribution [the act of attributing positive events and outcomes to one's own agency, but attributing negative events and outcomes to external forces]; and, (4) exaggeration [the act of representing outcomes or embellishing events as more significant than is actually suggested from other data].
Possible Limitations of the Researcher
- Access -- if your study depends on having access to people, organizations, data, or documents and, for whatever reason, access is denied or limited in some way, the reasons for this needs to be described. Also, include an explanation why being denied or limited access did not prevent you from following through on your study.
- Longitudinal effects -- unlike your professor, who can literally devote years [even a lifetime] to studying a single topic, the time available to investigate a research problem and to measure change or stability over time is constrained by the due date of your assignment. Be sure to choose a research problem that does not require an excessive amount of time to complete the literature review, apply the methodology, and gather and interpret the results. If you're unsure whether you can complete your research within the confines of the assignment's due date, talk to your professor.
- Cultural and other type of bias -- we all have biases, whether we are conscience of them or not. Bias is when a person, place, event, or thing is viewed or shown in a consistently inaccurate way. Bias is usually negative, though one can have a positive bias as well, especially if that bias reflects your reliance on research that only support your hypothesis. When proof-reading your paper, be especially critical in reviewing how you have stated a problem, selected the data to be studied, what may have been omitted, the manner in which you have ordered events, people, or places, how you have chosen to represent a person, place, or thing, to name a phenomenon, or to use possible words with a positive or negative connotation. NOTE : If you detect bias in prior research, it must be acknowledged and you should explain what measures were taken to avoid perpetuating that bias. For example, if a previous study only used boys to examine how music education supports effective math skills, describe how your research expands the study to include girls.
- Fluency in a language -- if your research focuses , for example, on measuring the perceived value of after-school tutoring among Mexican-American ESL [English as a Second Language] students and you are not fluent in Spanish, you are limited in being able to read and interpret Spanish language research studies on the topic or to speak with these students in their primary language. This deficiency should be acknowledged.
Aguinis, Hermam and Jeffrey R. Edwards. “Methodological Wishes for the Next Decade and How to Make Wishes Come True.” Journal of Management Studies 51 (January 2014): 143-174; Brutus, Stéphane et al. "Self-Reported Limitations and Future Directions in Scholarly Reports: Analysis and Recommendations." Journal of Management 39 (January 2013): 48-75; Senunyeme, Emmanuel K. Business Research Methods. Powerpoint Presentation. Regent University of Science and Technology; ter Riet, Gerben et al. “All That Glitters Isn't Gold: A Survey on Acknowledgment of Limitations in Biomedical Studies.” PLOS One 8 (November 2013): 1-6.
Structure and Writing Style
Information about the limitations of your study are generally placed either at the beginning of the discussion section of your paper so the reader knows and understands the limitations before reading the rest of your analysis of the findings, or, the limitations are outlined at the conclusion of the discussion section as an acknowledgement of the need for further study. Statements about a study's limitations should not be buried in the body [middle] of the discussion section unless a limitation is specific to something covered in that part of the paper. If this is the case, though, the limitation should be reiterated at the conclusion of the section.
If you determine that your study is seriously flawed due to important limitations , such as, an inability to acquire critical data, consider reframing it as an exploratory study intended to lay the groundwork for a more complete research study in the future. Be sure, though, to specifically explain the ways that these flaws can be successfully overcome in a new study.
But, do not use this as an excuse for not developing a thorough research paper! Review the tab in this guide for developing a research topic . If serious limitations exist, it generally indicates a likelihood that your research problem is too narrowly defined or that the issue or event under study is too recent and, thus, very little research has been written about it. If serious limitations do emerge, consult with your professor about possible ways to overcome them or how to revise your study.
When discussing the limitations of your research, be sure to:
- Describe each limitation in detailed but concise terms;
- Explain why each limitation exists;
- Provide the reasons why each limitation could not be overcome using the method(s) chosen to acquire or gather the data [cite to other studies that had similar problems when possible];
- Assess the impact of each limitation in relation to the overall findings and conclusions of your study; and,
- If appropriate, describe how these limitations could point to the need for further research.
Remember that the method you chose may be the source of a significant limitation that has emerged during your interpretation of the results [for example, you didn't interview a group of people that you later wish you had]. If this is the case, don't panic. Acknowledge it, and explain how applying a different or more robust methodology might address the research problem more effectively in a future study. A underlying goal of scholarly research is not only to show what works, but to demonstrate what doesn't work or what needs further clarification.
Aguinis, Hermam and Jeffrey R. Edwards. “Methodological Wishes for the Next Decade and How to Make Wishes Come True.” Journal of Management Studies 51 (January 2014): 143-174; Brutus, Stéphane et al. "Self-Reported Limitations and Future Directions in Scholarly Reports: Analysis and Recommendations." Journal of Management 39 (January 2013): 48-75; Ioannidis, John P.A. "Limitations are not Properly Acknowledged in the Scientific Literature." Journal of Clinical Epidemiology 60 (2007): 324-329; Pasek, Josh. Writing the Empirical Social Science Research Paper: A Guide for the Perplexed. January 24, 2012. Academia.edu; Structure: How to Structure the Research Limitations Section of Your Dissertation. Dissertations and Theses: An Online Textbook. Laerd.com; What Is an Academic Paper? Institute for Writing Rhetoric. Dartmouth College; Writing the Experimental Report: Methods, Results, and Discussion. The Writing Lab and The OWL. Purdue University.
Writing Tip
Don't Inflate the Importance of Your Findings!
After all the hard work and long hours devoted to writing your research paper, it is easy to get carried away with attributing unwarranted importance to what you’ve done. We all want our academic work to be viewed as excellent and worthy of a good grade, but it is important that you understand and openly acknowledge the limitations of your study. Inflating the importance of your study's findings could be perceived by your readers as an attempt hide its flaws or encourage a biased interpretation of the results. A small measure of humility goes a long way!
Another Writing Tip
Negative Results are Not a Limitation!
Negative evidence refers to findings that unexpectedly challenge rather than support your hypothesis. If you didn't get the results you anticipated, it may mean your hypothesis was incorrect and needs to be reformulated. Or, perhaps you have stumbled onto something unexpected that warrants further study. Moreover, the absence of an effect may be very telling in many situations, particularly in experimental research designs. In any case, your results may very well be of importance to others even though they did not support your hypothesis. Do not fall into the trap of thinking that results contrary to what you expected is a limitation to your study. If you carried out the research well, they are simply your results and only require additional interpretation.
Lewis, George H. and Jonathan F. Lewis. “The Dog in the Night-Time: Negative Evidence in Social Research.” The British Journal of Sociology 31 (December 1980): 544-558.
Yet Another Writing Tip
Sample Size Limitations in Qualitative Research
Sample sizes are typically smaller in qualitative research because, as the study goes on, acquiring more data does not necessarily lead to more information. This is because one occurrence of a piece of data, or a code, is all that is necessary to ensure that it becomes part of the analysis framework. However, it remains true that sample sizes that are too small cannot adequately support claims of having achieved valid conclusions and sample sizes that are too large do not permit the deep, naturalistic, and inductive analysis that defines qualitative inquiry. Determining adequate sample size in qualitative research is ultimately a matter of judgment and experience in evaluating the quality of the information collected against the uses to which it will be applied and the particular research method and purposeful sampling strategy employed. If the sample size is found to be a limitation, it may reflect your judgment about the methodological technique chosen [e.g., single life history study versus focus group interviews] rather than the number of respondents used.
Boddy, Clive Roland. "Sample Size for Qualitative Research." Qualitative Market Research: An International Journal 19 (2016): 426-432; Huberman, A. Michael and Matthew B. Miles. "Data Management and Analysis Methods." In Handbook of Qualitative Research . Norman K. Denzin and Yvonna S. Lincoln, eds. (Thousand Oaks, CA: Sage, 1994), pp. 428-444; Blaikie, Norman. "Confounding Issues Related to Determining Sample Size in Qualitative Research." International Journal of Social Research Methodology 21 (2018): 635-641; Oppong, Steward Harrison. "The Problem of Sampling in qualitative Research." Asian Journal of Management Sciences and Education 2 (2013): 202-210.
- << Previous: 8. The Discussion
- Next: 9. The Conclusion >>
- Last Updated: Jun 18, 2024 10:45 AM
- URL: https://libguides.usc.edu/writingguide
- Affiliate Program

- UNITED STATES
- 台灣 (TAIWAN)
- TÜRKIYE (TURKEY)
- Academic Editing Services
- - Research Paper
- - Journal Manuscript
- - Dissertation
- - College & University Assignments
- Admissions Editing Services
- - Application Essay
- - Personal Statement
- - Recommendation Letter
- - Cover Letter
- - CV/Resume
- Business Editing Services
- - Business Documents
- - Report & Brochure
- - Website & Blog
- Writer Editing Services
- - Script & Screenplay
- Our Editors
- Client Reviews
- Editing & Proofreading Prices
- Wordvice Points
- Partner Discount
- Plagiarism Checker
APA Citation Generator
MLA Citation Generator
Chicago Citation Generator
Vancouver Citation Generator
- - APA Style
- - MLA Style
- - Chicago Style
- - Vancouver Style
- Writing & Editing Guide
- Academic Resources
- Admissions Resources
How to Present the Limitations of the Study Examples
What are the limitations of a study?
The limitations of a study are the elements of methodology or study design that impact the interpretation of your research results. The limitations essentially detail any flaws or shortcomings in your study. Study limitations can exist due to constraints on research design, methodology, materials, etc., and these factors may impact the findings of your study. However, researchers are often reluctant to discuss the limitations of their study in their papers, feeling that bringing up limitations may undermine its research value in the eyes of readers and reviewers.
In spite of the impact it might have (and perhaps because of it) you should clearly acknowledge any limitations in your research paper in order to show readers—whether journal editors, other researchers, or the general public—that you are aware of these limitations and to explain how they affect the conclusions that can be drawn from the research.
In this article, we provide some guidelines for writing about research limitations, show examples of some frequently seen study limitations, and recommend techniques for presenting this information. And after you have finished drafting and have received manuscript editing for your work, you still might want to follow this up with academic editing before submitting your work to your target journal.
Why do I need to include limitations of research in my paper?
Although limitations address the potential weaknesses of a study, writing about them toward the end of your paper actually strengthens your study by identifying any problems before other researchers or reviewers find them.
Furthermore, pointing out study limitations shows that you’ve considered the impact of research weakness thoroughly and have an in-depth understanding of your research topic. Since all studies face limitations, being honest and detailing these limitations will impress researchers and reviewers more than ignoring them.

Where should I put the limitations of the study in my paper?
Some limitations might be evident to researchers before the start of the study, while others might become clear while you are conducting the research. Whether these limitations are anticipated or not, and whether they are due to research design or to methodology, they should be clearly identified and discussed in the discussion section —the final section of your paper. Most journals now require you to include a discussion of potential limitations of your work, and many journals now ask you to place this “limitations section” at the very end of your article.
Some journals ask you to also discuss the strengths of your work in this section, and some allow you to freely choose where to include that information in your discussion section—make sure to always check the author instructions of your target journal before you finalize a manuscript and submit it for peer review .
Limitations of the Study Examples
There are several reasons why limitations of research might exist. The two main categories of limitations are those that result from the methodology and those that result from issues with the researcher(s).
Common Methodological Limitations of Studies
Limitations of research due to methodological problems can be addressed by clearly and directly identifying the potential problem and suggesting ways in which this could have been addressed—and SHOULD be addressed in future studies. The following are some major potential methodological issues that can impact the conclusions researchers can draw from the research.
Issues with research samples and selection
Sampling errors occur when a probability sampling method is used to select a sample, but that sample does not reflect the general population or appropriate population concerned. This results in limitations of your study known as “sample bias” or “selection bias.”
For example, if you conducted a survey to obtain your research results, your samples (participants) were asked to respond to the survey questions. However, you might have had limited ability to gain access to the appropriate type or geographic scope of participants. In this case, the people who responded to your survey questions may not truly be a random sample.
Insufficient sample size for statistical measurements
When conducting a study, it is important to have a sufficient sample size in order to draw valid conclusions. The larger the sample, the more precise your results will be. If your sample size is too small, it will be difficult to identify significant relationships in the data.
Normally, statistical tests require a larger sample size to ensure that the sample is considered representative of a population and that the statistical result can be generalized to a larger population. It is a good idea to understand how to choose an appropriate sample size before you conduct your research by using scientific calculation tools—in fact, many journals now require such estimation to be included in every manuscript that is sent out for review.
Lack of previous research studies on the topic
Citing and referencing prior research studies constitutes the basis of the literature review for your thesis or study, and these prior studies provide the theoretical foundations for the research question you are investigating. However, depending on the scope of your research topic, prior research studies that are relevant to your thesis might be limited.
When there is very little or no prior research on a specific topic, you may need to develop an entirely new research typology. In this case, discovering a limitation can be considered an important opportunity to identify literature gaps and to present the need for further development in the area of study.
Methods/instruments/techniques used to collect the data
After you complete your analysis of the research findings (in the discussion section), you might realize that the manner in which you have collected the data or the ways in which you have measured variables has limited your ability to conduct a thorough analysis of the results.
For example, you might realize that you should have addressed your survey questions from another viable perspective, or that you were not able to include an important question in the survey. In these cases, you should acknowledge the deficiency or deficiencies by stating a need for future researchers to revise their specific methods for collecting data that includes these missing elements.
Common Limitations of the Researcher(s)
Study limitations that arise from situations relating to the researcher or researchers (whether the direct fault of the individuals or not) should also be addressed and dealt with, and remedies to decrease these limitations—both hypothetically in your study, and practically in future studies—should be proposed.
Limited access to data
If your research involved surveying certain people or organizations, you might have faced the problem of having limited access to these respondents. Due to this limited access, you might need to redesign or restructure your research in a different way. In this case, explain the reasons for limited access and be sure that your finding is still reliable and valid despite this limitation.
Time constraints
Just as students have deadlines to turn in their class papers, academic researchers might also have to meet deadlines for submitting a manuscript to a journal or face other time constraints related to their research (e.g., participants are only available during a certain period; funding runs out; collaborators move to a new institution). The time available to study a research problem and to measure change over time might be constrained by such practical issues. If time constraints negatively impacted your study in any way, acknowledge this impact by mentioning a need for a future study (e.g., a longitudinal study) to answer this research problem.
Conflicts arising from cultural bias and other personal issues
Researchers might hold biased views due to their cultural backgrounds or perspectives of certain phenomena, and this can affect a study’s legitimacy. Also, it is possible that researchers will have biases toward data and results that only support their hypotheses or arguments. In order to avoid these problems, the author(s) of a study should examine whether the way the research problem was stated and the data-gathering process was carried out appropriately.
Steps for Organizing Your Study Limitations Section
When you discuss the limitations of your study, don’t simply list and describe your limitations—explain how these limitations have influenced your research findings. There might be multiple limitations in your study, but you only need to point out and explain those that directly relate to and impact how you address your research questions.
We suggest that you divide your limitations section into three steps: (1) identify the study limitations; (2) explain how they impact your study in detail; and (3) propose a direction for future studies and present alternatives. By following this sequence when discussing your study’s limitations, you will be able to clearly demonstrate your study’s weakness without undermining the quality and integrity of your research.
Step 1. Identify the limitation(s) of the study
- This part should comprise around 10%-20% of your discussion of study limitations.
The first step is to identify the particular limitation(s) that affected your study. There are many possible limitations of research that can affect your study, but you don’t need to write a long review of all possible study limitations. A 200-500 word critique is an appropriate length for a research limitations section. In the beginning of this section, identify what limitations your study has faced and how important these limitations are.
You only need to identify limitations that had the greatest potential impact on: (1) the quality of your findings, and (2) your ability to answer your research question.

Step 2. Explain these study limitations in detail
- This part should comprise around 60-70% of your discussion of limitations.
After identifying your research limitations, it’s time to explain the nature of the limitations and how they potentially impacted your study. For example, when you conduct quantitative research, a lack of probability sampling is an important issue that you should mention. On the other hand, when you conduct qualitative research, the inability to generalize the research findings could be an issue that deserves mention.
Explain the role these limitations played on the results and implications of the research and justify the choice you made in using this “limiting” methodology or other action in your research. Also, make sure that these limitations didn’t undermine the quality of your dissertation .

Step 3. Propose a direction for future studies and present alternatives (optional)
- This part should comprise around 10-20% of your discussion of limitations.
After acknowledging the limitations of the research, you need to discuss some possible ways to overcome these limitations in future studies. One way to do this is to present alternative methodologies and ways to avoid issues with, or “fill in the gaps of” the limitations of this study you have presented. Discuss both the pros and cons of these alternatives and clearly explain why researchers should choose these approaches.
Make sure you are current on approaches used by prior studies and the impacts they have had on their findings. Cite review articles or scientific bodies that have recommended these approaches and why. This might be evidence in support of the approach you chose, or it might be the reason you consider your choices to be included as limitations. This process can act as a justification for your approach and a defense of your decision to take it while acknowledging the feasibility of other approaches.
P hrases and Tips for Introducing Your Study Limitations in the Discussion Section
The following phrases are frequently used to introduce the limitations of the study:
- “There may be some possible limitations in this study.”
- “The findings of this study have to be seen in light of some limitations.”
- “The first is the…The second limitation concerns the…”
- “The empirical results reported herein should be considered in the light of some limitations.”
- “This research, however, is subject to several limitations.”
- “The primary limitation to the generalization of these results is…”
- “Nonetheless, these results must be interpreted with caution and a number of limitations should be borne in mind.”
- “As with the majority of studies, the design of the current study is subject to limitations.”
- “There are two major limitations in this study that could be addressed in future research. First, the study focused on …. Second ….”
For more articles on research writing and the journal submissions and publication process, visit Wordvice’s Academic Resources page.
And be sure to receive professional English editing and proofreading services , including paper editing services , for your journal manuscript before submitting it to journal editors.
Wordvice Resources
Proofreading & Editing Guide
Writing the Results Section for a Research Paper
How to Write a Literature Review
Research Writing Tips: How to Draft a Powerful Discussion Section
How to Captivate Journal Readers with a Strong Introduction
Tips That Will Make Your Abstract a Success!
APA In-Text Citation Guide for Research Writing
Additional Resources
- Diving Deeper into Limitations and Delimitations (PhD student)
- Organizing Your Social Sciences Research Paper: Limitations of the Study (USC Library)
- Research Limitations (Research Methodology)
- How to Present Limitations and Alternatives (UMASS)
Article References
Pearson-Stuttard, J., Kypridemos, C., Collins, B., Mozaffarian, D., Huang, Y., Bandosz, P.,…Micha, R. (2018). Estimating the health and economic effects of the proposed US Food and Drug Administration voluntary sodium reformulation: Microsimulation cost-effectiveness analysis. PLOS. https://journals.plos.org/plosmedicine/article?id=10.1371/journal.pmed.1002551
Xu, W.L, Pedersen, N.L., Keller, L., Kalpouzos, G., Wang, H.X., Graff, C,. Fratiglioni, L. (2015). HHEX_23 AA Genotype Exacerbates Effect of Diabetes on Dementia and Alzheimer Disease: A Population-Based Longitudinal Study. PLOS. Retrieved from https://journals.plos.org/plosmedicine/article?id=10.1371/journal.pmed.1001853
How to present limitations in research
Last updated
30 January 2024
Reviewed by
Short on time? Get an AI generated summary of this article instead
Limitations don’t invalidate or diminish your results, but it’s best to acknowledge them. This will enable you to address any questions your study failed to answer because of them.
In this guide, learn how to recognize, present, and overcome limitations in research.
- What is a research limitation?
Research limitations are weaknesses in your research design or execution that may have impacted outcomes and conclusions. Uncovering limitations doesn’t necessarily indicate poor research design—it just means you encountered challenges you couldn’t have anticipated that limited your research efforts.
Does basic research have limitations?
Basic research aims to provide more information about your research topic . It requires the same standard research methodology and data collection efforts as any other research type, and it can also have limitations.
- Common research limitations
Researchers encounter common limitations when embarking on a study. Limitations can occur in relation to the methods you apply or the research process you design. They could also be connected to you as the researcher.
Methodology limitations
Not having access to data or reliable information can impact the methods used to facilitate your research. A lack of data or reliability may limit the parameters of your study area and the extent of your exploration.
Your sample size may also be affected because you won’t have any direction on how big or small it should be and who or what you should include. Having too few participants won’t adequately represent the population or groups of people needed to draw meaningful conclusions.
Research process limitations
The study’s design can impose constraints on the process. For example, as you’re conducting the research, issues may arise that don’t conform to the data collection methodology you developed. You may not realize until well into the process that you should have incorporated more specific questions or comprehensive experiments to generate the data you need to have confidence in your results.
Constraints on resources can also have an impact. Being limited on participants or participation incentives may limit your sample sizes. Insufficient tools, equipment, and materials to conduct a thorough study may also be a factor.
Common researcher limitations
Here are some of the common researcher limitations you may encounter:
Time: some research areas require multi-year longitudinal approaches, but you might not be able to dedicate that much time. Imagine you want to measure how much memory a person loses as they age. This may involve conducting multiple tests on a sample of participants over 20–30 years, which may be impossible.
Bias: researchers can consciously or unconsciously apply bias to their research. Biases can contribute to relying on research sources and methodologies that will only support your beliefs about the research you’re embarking on. You might also omit relevant issues or participants from the scope of your study because of your biases.
Limited access to data : you may need to pay to access specific databases or journals that would be helpful to your research process. You might also need to gain information from certain people or organizations but have limited access to them. These cases require readjusting your process and explaining why your findings are still reliable.
- Why is it important to identify limitations?
Identifying limitations adds credibility to research and provides a deeper understanding of how you arrived at your conclusions.
Constraints may have prevented you from collecting specific data or information you hoped would prove or disprove your hypothesis or provide a more comprehensive understanding of your research topic.
However, identifying the limitations contributing to your conclusions can inspire further research efforts that help gather more substantial information and data.
- Where to put limitations in a research paper
A research paper is broken up into different sections that appear in the following order:
Introduction
Methodology
The discussion portion of your paper explores your findings and puts them in the context of the overall research. Either place research limitations at the beginning of the discussion section before the analysis of your findings or at the end of the section to indicate that further research needs to be pursued.
What not to include in the limitations section
Evidence that doesn’t support your hypothesis is not a limitation, so you shouldn’t include it in the limitation section. Don’t just list limitations and their degree of severity without further explanation.
- How to present limitations
You’ll want to present the limitations of your study in a way that doesn’t diminish the validity of your research and leave the reader wondering if your results and conclusions have been compromised.
Include only the limitations that directly relate to and impact how you addressed your research questions. Following a specific format enables the reader to develop an understanding of the weaknesses within the context of your findings without doubting the quality and integrity of your research.
Identify the limitations specific to your study
You don’t have to identify every possible limitation that might have occurred during your research process. Only identify those that may have influenced the quality of your findings and your ability to answer your research question.
Explain study limitations in detail
This explanation should be the most significant portion of your limitation section.
Link each limitation with an interpretation and appraisal of their impact on the study. You’ll have to evaluate and explain whether the error, method, or validity issues influenced the study’s outcome and how.
Propose a direction for future studies and present alternatives
In this section, suggest how researchers can avoid the pitfalls you experienced during your research process.
If an issue with methodology was a limitation, propose alternate methods that may help with a smoother and more conclusive research project . Discuss the pros and cons of your alternate recommendation.
Describe steps taken to minimize each limitation
You probably took steps to try to address or mitigate limitations when you noticed them throughout the course of your research project. Describe these steps in the limitation section.
- Limitation example
“Approaches like stem cell transplantation and vaccination in AD [Alzheimer’s disease] work on a cellular or molecular level in the laboratory. However, translation into clinical settings will remain a challenge for the next decade.”
The authors are saying that even though these methods showed promise in helping people with memory loss when conducted in the lab (in other words, using animal studies), more studies are needed. These may be controlled clinical trials, for example.
However, the short life span of stem cells outside the lab and the vaccination’s severe inflammatory side effects are limitations. Researchers won’t be able to conduct clinical trials until these issues are overcome.
- How to overcome limitations in research
You’ve already started on the road to overcoming limitations in research by acknowledging that they exist. However, you need to ensure readers don’t mistake weaknesses for errors within your research design.
To do this, you’ll need to justify and explain your rationale for the methods, research design, and analysis tools you chose and how you noticed they may have presented limitations.
Your readers need to know that even when limitations presented themselves, you followed best practices and the ethical standards of your field. You didn’t violate any rules and regulations during your research process.
You’ll also want to reinforce the validity of your conclusions and results with multiple sources, methods, and perspectives. This prevents readers from assuming your findings were derived from a single or biased source.
- Learning and improving starts with limitations in research
Dealing with limitations with transparency and integrity helps identify areas for future improvements and developments. It’s a learning process, providing valuable insights into how you can improve methodologies, expand sample sizes, or explore alternate approaches to further support the validity of your findings.
Should you be using a customer insights hub?
Do you want to discover previous research faster?
Do you share your research findings with others?
Do you analyze research data?
Start for free today, add your research, and get to key insights faster
Editor’s picks
Last updated: 18 April 2023
Last updated: 27 February 2023
Last updated: 6 February 2023
Last updated: 15 January 2024
Last updated: 6 October 2023
Last updated: 5 February 2023
Last updated: 16 April 2023
Last updated: 7 March 2023
Last updated: 9 March 2023
Last updated: 12 December 2023
Last updated: 11 March 2024
Last updated: 13 May 2024
Latest articles
Related topics, .css-je19u9{-webkit-align-items:flex-end;-webkit-box-align:flex-end;-ms-flex-align:flex-end;align-items:flex-end;display:-webkit-box;display:-webkit-flex;display:-ms-flexbox;display:flex;-webkit-flex-direction:row;-ms-flex-direction:row;flex-direction:row;-webkit-box-flex-wrap:wrap;-webkit-flex-wrap:wrap;-ms-flex-wrap:wrap;flex-wrap:wrap;-webkit-box-pack:center;-ms-flex-pack:center;-webkit-justify-content:center;justify-content:center;row-gap:0;text-align:center;max-width:671px;}@media (max-width: 1079px){.css-je19u9{max-width:400px;}.css-je19u9>span{white-space:pre;}}@media (max-width: 799px){.css-je19u9{max-width:400px;}.css-je19u9>span{white-space:pre;}} decide what to .css-1kiodld{max-height:56px;display:-webkit-box;display:-webkit-flex;display:-ms-flexbox;display:flex;-webkit-align-items:center;-webkit-box-align:center;-ms-flex-align:center;align-items:center;}@media (max-width: 1079px){.css-1kiodld{display:none;}} build next, decide what to build next.

Users report unexpectedly high data usage, especially during streaming sessions.

Users find it hard to navigate from the home page to relevant playlists in the app.

It would be great to have a sleep timer feature, especially for bedtime listening.

I need better filters to find the songs or artists I’m looking for.
Log in or sign up
Get started for free
What are the limitations in research and how to write them?
Learn about the potential limitations in research and how to appropriately address them in order to deliver honest and ethical research.
It is fairly uncommon for researchers to stumble into the term research limitations when working on their research paper. Limitations in research can arise owing to constraints on design, methods, materials, and so on, and these aspects, unfortunately, may have an influence on your subject’s findings.
In this Mind The Graph’s article, we’ll discuss some recommendations for writing limitations in research , provide examples of various common types of limitations, and suggest how to properly present this information.
What are the limitations in research?
The limitations in research are the constraints in design, methods or even researchers’ limitations that affect and influence the interpretation of your research’s ultimate findings. These are limitations on the generalization and usability of findings that emerge from the design of the research and/or the method employed to ensure validity both internally and externally.
Researchers are usually cautious to acknowledge the limitations of their research in their publications for fear of undermining the research’s scientific validity. No research is faultless or covers every possible angle. As a result, addressing the constraints of your research exhibits honesty and integrity .
Why should include limitations of research in my paper?
Though limitations tackle potential flaws in research, commenting on them at the conclusion of your paper, by demonstrating that you are aware of these limitations and explaining how they impact the conclusions that may be taken from the research, improves your research by disclosing any issues before other researchers or reviewers do .
Additionally, emphasizing research constraints implies that you have thoroughly investigated the ramifications of research shortcomings and have a thorough understanding of your research problem.
Limits exist in any research; being honest about them and explaining them would impress researchers and reviewers more than disregarding them.
Remember that acknowledging a research’s shortcomings offers a chance to provide ideas for future research, but be careful to describe how your study may help to concentrate on these outstanding problems.
Possible limitations examples
Here are some limitations connected to methodology and the research procedure that you may need to explain and discuss in connection to your findings.
Methodological limitations
Sample size.
The number of units of analysis used in your study is determined by the sort of research issue being investigated. It is important to note that if your sample is too small, finding significant connections in the data will be challenging, as statistical tests typically require a larger sample size to ensure a fair representation and this can be limiting.
Lack of available or reliable data
A lack of data or trustworthy data will almost certainly necessitate limiting the scope of your research or the size of your sample, or it can be a substantial impediment to identifying a pattern and a relevant connection.
Lack of prior research on the subject
Citing previous research papers forms the basis of your literature review and aids in comprehending the research subject you are researching. Yet there may be little if any, past research on your issue.
The measure used to collect data
After finishing your analysis of the findings, you realize that the method you used to collect data limited your capacity to undertake a comprehensive evaluation of the findings. Recognize the flaw by mentioning that future researchers should change the specific approach for data collection.
Issues with research samples and selection
Sampling inaccuracies arise when a probability sampling method is employed to choose a sample, but that sample does not accurately represent the overall population or the relevant group. As a result, your study suffers from “sampling bias” or “selection bias.”
Limitations of the research
When your research requires polling certain persons or a specific group, you may have encountered the issue of limited access to these interviewees. Because of the limited access, you may need to reorganize or rearrange your research. In this scenario, explain why access is restricted and ensure that your findings are still trustworthy and valid despite the constraint.
Time constraints
Practical difficulties may limit the amount of time available to explore a research issue and monitor changes as they occur. If time restrictions have any detrimental influence on your research, recognize this impact by expressing the necessity for a future investigation.
Due to their cultural origins or opinions on observed events, researchers may carry biased opinions, which can influence the credibility of a research. Furthermore, researchers may exhibit biases toward data and conclusions that only support their hypotheses or arguments.
The structure of the limitations section
The limitations of your research are usually stated at the beginning of the discussion section of your paper so that the reader is aware of and comprehends the limitations prior to actually reading the rest of your findings, or they are stated at the end of the discussion section as an acknowledgment of the need for further research.
The ideal way is to divide your limitations section into three steps:
1. Identify the research constraints;
2. Describe in great detail how they affect your research;
3. Mention the opportunity for future investigations and give possibilities.
By following this method while addressing the constraints of your research, you will be able to effectively highlight your research’s shortcomings without jeopardizing the quality and integrity of your research.
Present your research or paper in an innovative way
If you want your readers to be engaged and participate in your research, try Mind The Graph tool to add visual assets to your content. Infographics may improve comprehension and are easy to read, just as the Mind The Graph tool is simple to use and offers a variety of templates from which you can select the one that best suits your information.
Subscribe to our newsletter
Exclusive high quality content about effective visual communication in science.
About Jessica Abbadia
Jessica Abbadia is a lawyer that has been working in Digital Marketing since 2020, improving organic performance for apps and websites in various regions through ASO and SEO. Currently developing scientific and intellectual knowledge for the community's benefit. Jessica is an animal rights activist who enjoys reading and drinking strong coffee.
Content tags

Writing Limitations of Research Study — 4 Reasons Why It Is Important!
It is not unusual for researchers to come across the term limitations of research during their academic paper writing. More often this is interpreted as something terrible. However, when it comes to research study, limitations can help structure the research study better. Therefore, do not underestimate significance of limitations of research study.
Allow us to take you through the context of how to evaluate the limits of your research and conclude an impactful relevance to your results.
Table of Contents
What Are the Limitations of a Research Study?
Every research has its limit and these limitations arise due to restrictions in methodology or research design. This could impact your entire research or the research paper you wish to publish. Unfortunately, most researchers choose not to discuss their limitations of research fearing it will affect the value of their article in the eyes of readers.
However, it is very important to discuss your study limitations and show it to your target audience (other researchers, journal editors, peer reviewers etc.). It is very important that you provide an explanation of how your research limitations may affect the conclusions and opinions drawn from your research. Moreover, when as an author you state the limitations of research, it shows that you have investigated all the weaknesses of your study and have a deep understanding of the subject. Being honest could impress your readers and mark your study as a sincere effort in research.

Why and Where Should You Include the Research Limitations?
The main goal of your research is to address your research objectives. Conduct experiments, get results and explain those results, and finally justify your research question . It is best to mention the limitations of research in the discussion paragraph of your research article.
At the very beginning of this paragraph, immediately after highlighting the strengths of the research methodology, you should write down your limitations. You can discuss specific points from your research limitations as suggestions for further research in the conclusion of your thesis.
1. Common Limitations of the Researchers
Limitations that are related to the researcher must be mentioned. This will help you gain transparency with your readers. Furthermore, you could provide suggestions on decreasing these limitations in you and your future studies.
2. Limited Access to Information
Your work may involve some institutions and individuals in research, and sometimes you may have problems accessing these institutions. Therefore, you need to redesign and rewrite your work. You must explain your readers the reason for limited access.
3. Limited Time
All researchers are bound by their deadlines when it comes to completing their studies. Sometimes, time constraints can affect your research negatively. However, the best practice is to acknowledge it and mention a requirement for future study to solve the research problem in a better way.
4. Conflict over Biased Views and Personal Issues
Biased views can affect the research. In fact, researchers end up choosing only those results and data that support their main argument, keeping aside the other loose ends of the research.
Types of Limitations of Research
Before beginning your research study, know that there are certain limitations to what you are testing or possible research results. There are different types that researchers may encounter, and they all have unique characteristics, such as:
1. Research Design Limitations
Certain restrictions on your research or available procedures may affect your final results or research outputs. You may have formulated research goals and objectives too broadly. However, this can help you understand how you can narrow down the formulation of research goals and objectives, thereby increasing the focus of your study.
2. Impact Limitations
Even if your research has excellent statistics and a strong design, it can suffer from the influence of the following factors:
- Presence of increasing findings as researched
- Being population specific
- A strong regional focus.
3. Data or statistical limitations
In some cases, it is impossible to collect sufficient data for research or very difficult to get access to the data. This could lead to incomplete conclusion to your study. Moreover, this insufficiency in data could be the outcome of your study design. The unclear, shabby research outline could produce more problems in interpreting your findings.
How to Correctly Structure Your Research Limitations?
There are strict guidelines for narrowing down research questions, wherein you could justify and explain potential weaknesses of your academic paper. You could go through these basic steps to get a well-structured clarity of research limitations:
- Declare that you wish to identify your limitations of research and explain their importance,
- Provide the necessary depth, explain their nature, and justify your study choices.
- Write how you are suggesting that it is possible to overcome them in the future.
In this section, your readers will see that you are aware of the potential weaknesses in your business, understand them and offer effective solutions, and it will positively strengthen your article as you clarify all limitations of research to your target audience.
Know that you cannot be perfect and there is no individual without flaws. You could use the limitations of research as a great opportunity to take on a new challenge and improve the future of research. In a typical academic paper, research limitations may relate to:
1. Formulating your goals and objectives
If you formulate goals and objectives too broadly, your work will have some shortcomings. In this case, specify effective methods or ways to narrow down the formula of goals and aim to increase your level of study focus.
2. Application of your data collection methods in research
If you do not have experience in primary data collection, there is a risk that there will be flaws in the implementation of your methods. It is necessary to accept this, and learn and educate yourself to understand data collection methods.
3. Sample sizes
This depends on the nature of problem you choose. Sample size is of a greater importance in quantitative studies as opposed to qualitative ones. If your sample size is too small, statistical tests cannot identify significant relationships or connections within a given data set.
You could point out that other researchers should base the same study on a larger sample size to get more accurate results.
4. The absence of previous studies in the field you have chosen
Writing a literature review is an important step in any scientific study because it helps researchers determine the scope of current work in the chosen field. It is a major foundation for any researcher who must use them to achieve a set of specific goals or objectives.
However, if you are focused on the most current and evolving research problem or a very narrow research problem, there may be very little prior research on your topic. For example, if you chose to explore the role of Bitcoin as the currency of the future, you may not find tons of scientific papers addressing the research problem as Bitcoins are only a new phenomenon.
It is important that you learn to identify research limitations examples at each step. Whatever field you choose, feel free to add the shortcoming of your work. This is mainly because you do not have many years of experience writing scientific papers or completing complex work. Therefore, the depth and scope of your discussions may be compromised at different levels compared to academics with a lot of expertise. Include specific points from limitations of research. Use them as suggestions for the future.
Have you ever faced a challenge of writing the limitations of research study in your paper? How did you overcome it? What ways did you follow? Were they beneficial? Let us know in the comments below!
Frequently Asked Questions
Setting limitations in our study helps to clarify the outcomes drawn from our research and enhance understanding of the subject. Moreover, it shows that the author has investigated all the weaknesses in the study.
Scope is the range and limitations of a research project which are set to define the boundaries of a project. Limitations are the impacts on the overall study due to the constraints on the research design.
Limitation in research is an impact of a constraint on the research design in the overall study. They are the flaws or weaknesses in the study, which may influence the outcome of the research.
1. Limitations in research can be written as follows: Formulate your goals and objectives 2. Analyze the chosen data collection method and the sample sizes 3. Identify your limitations of research and explain their importance 4. Provide the necessary depth, explain their nature, and justify your study choices 5. Write how you are suggesting that it is possible to overcome them in the future
Excellent article ,,,it has helped me big
This is very helpful information. It has given me an insight on how to go about my study limitations.
Good comments and helpful
the topic is well covered
Rate this article Cancel Reply
Your email address will not be published.

Enago Academy's Most Popular Articles

- Publishing Research
- Trending Now
- Understanding Ethics
Understanding the Impact of Retractions on Research Integrity – A global study
As we reach the midway point of 2024, ‘Research Integrity’ remains one of the hot…

- Diversity and Inclusion
The Silent Struggle: Confronting gender bias in science funding
In the 1990s, Dr. Katalin Kariko’s pioneering mRNA research seemed destined for obscurity, doomed by…

- Promoting Research
Plain Language Summary — Communicating your research to bridge the academic-lay gap
Science can be complex, but does that mean it should not be accessible to the…


Addressing Barriers in Academia: Navigating unconscious biases in the Ph.D. journey
In the journey of academia, a Ph.D. marks a transitional phase, like that of a…

- Manuscripts & Grants
- Reporting Research
Unraveling Research Population and Sample: Understanding their role in statistical inference
Research population and sample serve as the cornerstones of any scientific inquiry. They hold the…
Research Problem Statement — Find out how to write an impactful one!
How to Develop a Good Research Question? — Types & Examples
5 Effective Ways to Avoid Ghostwriting for Busy Researchers

Sign-up to read more
Subscribe for free to get unrestricted access to all our resources on research writing and academic publishing including:
- 2000+ blog articles
- 50+ Webinars
- 10+ Expert podcasts
- 50+ Infographics
- 10+ Checklists
- Research Guides
We hate spam too. We promise to protect your privacy and never spam you.

Stating the Obvious: Writing Assumptions, Limitations, and Delimitations

During the process of writing your thesis or dissertation, you might suddenly realize that your research has inherent flaws. Don’t worry! Virtually all projects contain restrictions to your research. However, being able to recognize and accurately describe these problems is the difference between a true researcher and a grade-school kid with a science-fair project. Concerns with truthful responding, access to participants, and survey instruments are just a few of examples of restrictions on your research. In the following sections, the differences among delimitations, limitations, and assumptions of a dissertation will be clarified.
Delimitations
Delimitations are the definitions you set as the boundaries of your own thesis or dissertation, so delimitations are in your control. Delimitations are set so that your goals do not become impossibly large to complete. Examples of delimitations include objectives, research questions, variables, theoretical objectives that you have adopted, and populations chosen as targets to study. When you are stating your delimitations, clearly inform readers why you chose this course of study. The answer might simply be that you were curious about the topic and/or wanted to improve standards of a professional field by revealing certain findings. In any case, you should clearly list the other options available and the reasons why you did not choose these options immediately after you list your delimitations. You might have avoided these options for reasons of practicality, interest, or relativity to the study at hand. For example, you might have only studied Hispanic mothers because they have the highest rate of obese babies. Delimitations are often strongly related to your theory and research questions. If you were researching whether there are different parenting styles between unmarried Asian, Caucasian, African American, and Hispanic women, then a delimitation of your study would be the inclusion of only participants with those demographics and the exclusion of participants from other demographics such as men, married women, and all other ethnicities of single women (inclusion and exclusion criteria). A further delimitation might be that you only included closed-ended Likert scale responses in the survey, rather than including additional open-ended responses, which might make some people more willing to take and complete your survey. Remember that delimitations are not good or bad. They are simply a detailed description of the scope of interest for your study as it relates to the research design. Don’t forget to describe the philosophical framework you used throughout your study, which also delimits your study.
Limitations
Limitations of a dissertation are potential weaknesses in your study that are mostly out of your control, given limited funding, choice of research design, statistical model constraints, or other factors. In addition, a limitation is a restriction on your study that cannot be reasonably dismissed and can affect your design and results. Do not worry about limitations because limitations affect virtually all research projects, as well as most things in life. Even when you are going to your favorite restaurant, you are limited by the menu choices. If you went to a restaurant that had a menu that you were craving, you might not receive the service, price, or location that makes you enjoy your favorite restaurant. If you studied participants’ responses to a survey, you might be limited in your abilities to gain the exact type or geographic scope of participants you wanted. The people whom you managed to get to take your survey may not truly be a random sample, which is also a limitation. If you used a common test for data findings, your results are limited by the reliability of the test. If your study was limited to a certain amount of time, your results are affected by the operations of society during that time period (e.g., economy, social trends). It is important for you to remember that limitations of a dissertation are often not something that can be solved by the researcher. Also, remember that whatever limits you also limits other researchers, whether they are the largest medical research companies or consumer habits corporations. Certain kinds of limitations are often associated with the analytical approach you take in your research, too. For example, some qualitative methods like heuristics or phenomenology do not lend themselves well to replicability. Also, most of the commonly used quantitative statistical models can only determine correlation, but not causation.
Assumptions
Assumptions are things that are accepted as true, or at least plausible, by researchers and peers who will read your dissertation or thesis. In other words, any scholar reading your paper will assume that certain aspects of your study is true given your population, statistical test, research design, or other delimitations. For example, if you tell your friend that your favorite restaurant is an Italian place, your friend will assume that you don’t go there for the sushi. It’s assumed that you go there to eat Italian food. Because most assumptions are not discussed in-text, assumptions that are discussed in-text are discussed in the context of the limitations of your study, which is typically in the discussion section. This is important, because both assumptions and limitations affect the inferences you can draw from your study. One of the more common assumptions made in survey research is the assumption of honesty and truthful responses. However, for certain sensitive questions this assumption may be more difficult to accept, in which case it would be described as a limitation of the study. For example, asking people to report their criminal behavior in a survey may not be as reliable as asking people to report their eating habits. It is important to remember that your limitations and assumptions should not contradict one another. For instance, if you state that generalizability is a limitation of your study given that your sample was limited to one city in the United States, then you should not claim generalizability to the United States population as an assumption of your study. Statistical models in quantitative research designs are accompanied with assumptions as well, some more strict than others. These assumptions generally refer to the characteristics of the data, such as distributions, correlational trends, and variable type, just to name a few. Violating these assumptions can lead to drastically invalid results, though this often depends on sample size and other considerations.
Click here to cancel reply.
You must be logged in to post a comment.
Copyright © 2024 PhDStudent.com. All rights reserved. Designed by Divergent Web Solutions, LLC .
- The Scientist University
How to Present a Research Study’s Limitations
All studies have imperfections, but how to present them without diminishing the value of the work can be tricky..

Nathan Ni holds a PhD from Queens University. He is a science editor for The Scientist’s Creative Services Team who strives to better understand and communicate the relationships between health and disease.
View full profile.
Learn about our editorial policies.

Scientists work with many different limitations. First and foremost, they navigate informational limitations, work around knowledge gaps when designing studies, formulating hypotheses, and analyzing data. They also handle technical limitations, making the most of what their hands, equipment, and instruments can achieve. Finally, researchers must also manage logistical limitations. Scientists will often experience sample scarcity, financial issues, or simply be unable to access the technology or materials that they want.
All scientific studies have limitations, and no study is perfect. Researchers should not run from this reality, but engage it directly. It is better to directly address the specific limitations of the work in question, and doing so is actually a way to demonstrate an author’s proficiency and aptitude.
Do: Be Transparent
From a practical perspective, being transparent is the main key to directly addressing the specific limitations of a study. Was there an experiment that the researchers wanted to perform but could not, or a sample that existed that the scientists could not obtain? Was there a piece of knowledge that would explain a question raised by the data presented within the current study? If the answer is yes, the authors should mention this and elaborate upon it within the discussion section.
Asking and addressing these questions demonstrates that the authors have knowledge, understanding, and expertise of the subject area beyond what the study directly investigated. It further demonstrates a solid grasp of the existing literature—which means a solid grasp of what others are doing, what techniques they are using, and what limitations impede their own studies. This information helps the authors contextualize where their study fits within what others have discovered, thereby mitigating the perceived effect of a given limitation on the study’s legitimacy. In essence, this strategy turns limitations, often considered weaknesses, into strengths.
For example, in their 2021 Cell Reports study on macrophage polarization mechanisms, dermatologist Alexander Marneros and colleagues wrote the following. 1
A limitation of studying macrophage polarization in vitro is that this approach only partially captures the tissue microenvironment context in which many different factors affect macrophage polarization. However, it is likely that the identified signaling mechanisms that promote polarization in vitro are also critical for polarization mechanisms that occur in vivo. This is supported by our observation that trametinib and panobinostat inhibited M2-type macrophage polarization not only in vitro but also in skin wounds and laser-induced CNV lesions.
This is a very effective structure. In the first sentence ( yellow ), the authors outlined the limitation. In the next sentence ( green ), they offered a rationalization that mitigates the effect of the limitation. Finally, they provided the evidence ( blue ) for this rationalization, using not just information from the literature, but also data that they obtained in their study specifically for this purpose.

Don't: Be Defensive
It can feel natural to avoid talking about a study’s limitations. Scientists may believe that mentioning the drawbacks still present in their study will jeopardize their chances of publication. As such, researchers will sometimes skirt around the issue. They will present “boilerplate faults”—generalized concerns about sample size/diversity and time limitations that all researchers face—rather than honestly discussing their own study. Alternatively, they will describe their limitations in a defensive manner, positioning their problems as something that “could not be helped”—as something beyond what science can currently achieve.
However, their audience can see through this, because they are largely peers who understand and have experienced how modern research works. They can tell the difference between global challenges faced by every scientific study and limitations that are specific to a single study. Avoiding these specific limitations can therefore betray a lack of confidence that the study is good enough to withstand problems stemming from legitimate limitations. As such, researchers should actively engage with the greater scientific implications of the limitations that they face. Indeed, doing this is actually a way to demonstrate an author’s proficiency and aptitude.
In an example, neurogeneticist Nancy Bonini and colleagues, in their publication in Nature , discussed a question raised by their data that they have elected not to directly investigate in this study, writing “ Among the intriguing questions raised by these data is how senescent glia promote LDs in other glia. ” To show both the legitimacy of the question and how seriously they have considered it, the authors provided a comprehensive summary of the literature in the following seven sentences, offering two hypotheses backed by a combined eight different sources. 2 Rather than shying away from a limitation, they attacked it as something to be curious about and to discuss. This is not just a very effective way of demonstrating their expertise, but it frames the limitation as something that, when overcome, will build upon the present study rather than something that negatively affects the legitimacy of their current findings.
Striking the Right Balance
Scientists have to navigate the fine line between acknowledging the limitations of their study while also not diminishing the effect and value of their own work. To be aware of legitimate limitations and properly assess and dissect them shows a profound understanding of a field, where the study fits within that field, and what the rest of the scientific community are doing and what challenges they face.
All studies are parts of a greater whole. Pretending otherwise is a disservice to the scientific community.
Looking for more information on scientific writing? Check out The Scientist’ s TS SciComm section. Looking for some help putting together a manuscript, a figure, a poster, or anything else? The Scientist ’s Scientific Services may have the professional help that you need.
- He L, et al. Global characterization of macrophage polarization mechanisms and identification of M2-type polarization inhibitors . Cell Rep . 2021;37(5):109955.
- Byrns CN, et al. Senescent glia link mitochondrial dysfunction and lipid accumulation . Nature . 2024.
Training videos | Faqs

Limitations in Research – A Simplified Guide with Examples
Limitations are flaws and shortcomings of your study. It is very important that you discuss the limitations of your study in the discussion section of your research paper. In this blog, we provide tips for presenting study limitations in your paper along with some real-world examples.
1. Should I Report the Limitations of My Work?

Most studies will have some form of limitation. So be honest and don’t hide your limitations. You have to tell your readers how your limitations might influence the outcomes and conclusions of your research. In reality, your readers and reviewers will be impressed with your paper if you are upfront about your limitations.
2. Examples
Let’s look at some examples. We have selected a variety of examples from different research topics.
2.1. Limitations Example 1
Following example is from a Medical research paper.
✔ The authors talk about the limitations and emphasis the importance of reconfirming the findings in a much larger study Study design and small sample size are important limitations. This could have led to an overestimation of the effect. Future research should reconfirm these findings by conducting larger-scale studies. _ Limitation s _ How it might affect the results? _ How to fix the limitation?
The authors are saying that the main limitations of the study are the small sample size and weak study design. Then they explain how this might have affected their results. They are saying that it is possible that they are overestimating the actual effect they are measuring. Then finally they are telling the readers that more studies with larger sample sizes should be conducted to reconfirm the findings.
As you can see, the authors are clearly explaining three things here: (1) What is the limitation? (2) How it might affect the study outcomes? and (3) What should be done to address the limitation?
2.2. Limitations Example 2
Following example is from an Engineering research paper.
✔ The authors are acknowledging the limitations and warning readers against generalizing the research findings However, some study limitations should be acknowledged. The experiments do not fully consider the problems that can appear in real situations. Hence, caution should be taken with generalizing the findings and applying them to real-life situations. _ Acknowledging limitations _ Explaining the limitation _ How it might affect the results?
The authors acknowledge that their study has some limitations. Then they explain what the limitations are. They are saying that their experiments do not consider all problems that might occur in real-life situations. Then they explain how this might affect their research outcomes. They are saying that readers should be careful when generalizing the results to practical real-world situations, because there is a possibility that the methods might fail.
2.3. Limitations Example 3
It is important to remember not to end your paper with limitations. Finish your paper on a positive note by telling your readers about the benefits of your research and possible future directions. In the following example, right after listing the limitations, the authors proceed to talk about the positive aspects of the work.
✔ The authors finish their paper on a positive note by talking about the benefits of their work and possible future work With this limited study, it is not known whether this finding can be applied to all clinical scenarios. Notwithstanding these limitations, this study has proven that Ultrasound can potentially serve as a more efficient alternative to X-rays in diagnosis. Future directions include studying the effects of different ultrasound pulsing schemes on pain relief. Another interesting direction would be to consider applications in nonhuman primates. _ Limitations _ Benefits of the work _ Possible future directions
The authors are saying that their experiments were somewhat limited and are not sure if their findings apply to the wider clinical practice. Then the authors highlight the benefits of their research. The authors say that their study has proven that ultrasound can be used instead of X-rays for diagnosis of certain types of diseases. Then they are explaining how future research can extend this work further. The authors are suggesting that it will be interesting to explore if ultrasound can be used for the treatment of chronic pain. And they are also suggesting that future studies can explore treating certain types of animal diseases with ultrasound. This is a very good example of how to finish the discussion section of your paper on a positive note.
Limitations are a vital component of the discussion section of your research paper. Remember, every study has limitations. There is no such thing as a perfect study. One of the major mistakes beginner writers make is hiding the limitations in the paper. Don’t do this, reviewers will reject your paper. Explain clearly how your limitations might have impacted your results, and provide ideas to mitigate them in the future. For further reading, please refer to our blogs on handling negative results and advanced tactics to address study limitations.
If you have any questions, please drop a comment below, and we will answer as soon as possible. We also recommend you to refer to our other blogs on academic writing tools , academic writing resources , academic writing phrases and research paper examples which are relevant to the topic discussed in this blog.
Similar Posts

Formulating Strong Research Questions: Examples and Writing Tips
In this blog, we will go through many research question examples and understand how to construct a strong research question for your paper.

Useful Phrases and Sentences for Academic & Research Paper Writing
In this blog, we explain various sections of a research paper and give you an overview of what these sections should contain.

Mastering Verb Tenses in Literature Reviews
In this blog, we will see what tense you should use in the literature review section of your research paper.

Literature Review Examples and Writing Tips
In this blog, we will go through many literature review examples and understand different ways to present past literature in your paper.

Writing a Medical Clinical Trial Research Paper – Example & Format
In this blog, we will teach you step-by-step how to write a clinical trial research paper for publication in a high quality scientific journal.

How to Make Your Study Limitations Sound Positive?
In this blog, we will look at some clever techniques to present the study limitations without reducing the impact of your work.
Leave a Reply Cancel reply
Your email address will not be published. Required fields are marked *
Save my name, email, and website in this browser for the next time I comment.
- 10 Share Facebook
- 1 Share Twitter
- 0 Share LinkedIn
- 0 Share Email

Organizing Academic Research Papers: Limitations of the Study
- Purpose of Guide
- Design Flaws to Avoid
- Glossary of Research Terms
- Narrowing a Topic Idea
- Broadening a Topic Idea
- Extending the Timeliness of a Topic Idea
- Academic Writing Style
- Choosing a Title
- Making an Outline
- Paragraph Development
- Executive Summary
- Background Information
- The Research Problem/Question
- Theoretical Framework
- Citation Tracking
- Content Alert Services
- Evaluating Sources
- Primary Sources
- Secondary Sources
- Tertiary Sources
- What Is Scholarly vs. Popular?
- Qualitative Methods
- Quantitative Methods
- Using Non-Textual Elements
- Limitations of the Study
- Common Grammar Mistakes
- Avoiding Plagiarism
- Footnotes or Endnotes?
- Further Readings
- Annotated Bibliography
- Dealing with Nervousness
- Using Visual Aids
- Grading Someone Else's Paper
- How to Manage Group Projects
- Multiple Book Review Essay
- Reviewing Collected Essays
- About Informed Consent
- Writing Field Notes
- Writing a Policy Memo
- Writing a Research Proposal
- Acknowledgements
The limitations of the study are those characteristics of design or methodology that impacted or influenced the application or interpretation of the results of your study. They are the constraints on generalizability and utility of findings that are the result of the ways in which you chose to design the study and/or the method used to establish internal and external validity.
Importance of...
Always acknowledge a study's limitations. It is far better for you to identify and acknowledge your study’s limitations than to have them pointed out by your professor and be graded down because you appear to have ignored them.
Keep in mind that acknowledgement of a study's limitations is an opportunity to make suggestions for further research. If you do connect your study's limitations to suggestions for further research, be sure to explain the ways in which these unanswered questions may become more focused because of your study.
Acknowledgement of a study's limitations also provides you with an opportunity to demonstrate to your professor that you have thought critically about the research problem, understood the relevant literature published about it, and correctly assessed the methods chosen for studying the problem. A key objective of the research process is not only discovering new knowledge but also to confront assumptions and explore what we don't know.
Claiming limitiations is a subjective process because you must evaluate the impact of those limitations . Don't just list key weaknesses and the magnitude of a study's limitations. To do so diminishes the validity of your research because it leaves the reader wondering whether, or in what ways, limitation(s) in your study may have impacted the findings and conclusions. Limitations require a critical, overall appraisal and interpretation of their impact. You should answer the question: do these problems with errors, methods, validity, etc. eventually matter and, if so, to what extent?
Structure: How to Structure the Research Limitations Section of Your Dissertation . Dissertations and Theses: An Online Textbook. Laerd.com.
Descriptions of Possible Limitations
All studies have limitations . However, it is important that you restrict your discussion to limitations related to the research problem under investigation. For example, if a meta-analysis of existing literature is not a stated purpose of your research, it should not be discussed as a limitation. Do not apologize for not addressing issues that you did not promise to investigate in your paper.
Here are examples of limitations you may need to describe and to discuss how they possibly impacted your findings. Descriptions of limitations should be stated in the past tense.
Possible Methodological Limitations
- Sample size -- the number of the units of analysis you use in your study is dictated by the type of research problem you are investigating. Note that, if your sample size is too small, it will be difficult to find significant relationships from the data, as statistical tests normally require a larger sample size to ensure a representative distribution of the population and to be considered representative of groups of people to whom results will be generalized or transferred.
- Lack of available and/or reliable data -- a lack of data or of reliable data will likely require you to limit the scope of your analysis, the size of your sample, or it can be a significant obstacle in finding a trend and a meaningful relationship. You need to not only describe these limitations but to offer reasons why you believe data is missing or is unreliable. However, don’t just throw up your hands in frustration; use this as an opportunity to describe the need for future research.
- Lack of prior research studies on the topic -- citing prior research studies forms the basis of your literature review and helps lay a foundation for understanding the research problem you are investigating. Depending on the currency or scope of your research topic, there may be little, if any, prior research on your topic. Before assuming this to be true, consult with a librarian! In cases when a librarian has confirmed that there is a lack of prior research, you may be required to develop an entirely new research typology [for example, using an exploratory rather than an explanatory research design]. Note that this limitation can serve as an important opportunity to describe the need for further research.
- Measure used to collect the data -- sometimes it is the case that, after completing your interpretation of the findings, you discover that the way in which you gathered data inhibited your ability to conduct a thorough analysis of the results. For example, you regret not including a specific question in a survey that, in retrospect, could have helped address a particular issue that emerged later in the study. Acknowledge the deficiency by stating a need in future research to revise the specific method for gathering data.
- Self-reported data -- whether you are relying on pre-existing self-reported data or you are conducting a qualitative research study and gathering the data yourself, self-reported data is limited by the fact that it rarely can be independently verified. In other words, you have to take what people say, whether in interviews, focus groups, or on questionnaires, at face value. However, self-reported data contain several potential sources of bias that should be noted as limitations: (1) selective memory (remembering or not remembering experiences or events that occurred at some point in the past); (2) telescoping [recalling events that occurred at one time as if they occurred at another time]; (3) attribution [the act of attributing positive events and outcomes to one's own agency but attributing negative events and outcomes to external forces]; and, (4) exaggeration [the act of representing outcomes or embellishing events as more significant than is actually suggested from other data].
Possible Limitations of the Researcher
- Access -- if your study depends on having access to people, organizations, or documents and, for whatever reason, access is denied or otherwise limited, the reasons for this need to be described.
- Longitudinal effects -- unlike your professor, who can literally devote years [even a lifetime] to studying a single research problem, the time available to investigate a research problem and to measure change or stability within a sample is constrained by the due date of your assignment. Be sure to choose a topic that does not require an excessive amount of time to complete the literature review, apply the methodology, and gather and interpret the results. If you're unsure, talk to your professor.
- Cultural and other type of bias -- we all have biases, whether we are conscience of them or not. Bias is when a person, place, or thing is viewed or shown in a consistently inaccurate way. It is usually negative, though one can have a positive bias as well. When proof-reading your paper, be especially critical in reviewing how you have stated a problem, selected the data to be studied, what may have been omitted, the manner in which you have ordered events, people, or places and how you have chosen to represent a person, place, or thing, to name a phenomenon, or to use possible words with a positive or negative connotation. Note that if you detect bias in prior research, it must be acknowledged and you should explain what measures were taken to avoid perpetuating bias.
- Fluency in a language -- if your research focuses on measuring the perceived value of after-school tutoring among Mexican-American ESL [English as a Second Language] students, for example, and you are not fluent in Spanish, you are limited in being able to read and interpret Spanish language research studies on the topic. This deficiency should be acknowledged.
Brutus, Stéphane et al. Self-Reported Limitations and Future Directions in Scholarly Reports: Analysis and Recommendations. Journal of Management 39 (January 2013): 48-75; Senunyeme, Emmanuel K. Business Research Methods . Powerpoint Presentation. Regent University of Science and Technology.
Structure and Writing Style
Information about the limitations of your study are generally placed either at the beginning of the discussion section of your paper so the reader knows and understands the limitations before reading the rest of your analysis of the findings, or, the limitations are outlined at the conclusion of the discussion section as an acknowledgement of the need for further study. Statements about a study's limitations should not be buried in the body [middle] of the discussion section unless a limitation is specific to something covered in that part of the paper. If this is the case, though, the limitation should be reiterated at the conclusion of the section.
If you determine that your study is seriously flawed due to important limitations , such as, an inability to acquire critical data, consider reframing it as a pilot study intended to lay the groundwork for a more complete research study in the future. Be sure, though, to specifically explain the ways that these flaws can be successfully overcome in later studies.
But, do not use this as an excuse for not developing a thorough research paper! Review the tab in this guide for developing a research topic . If serious limitations exist, it generally indicates a likelihood that your research problem is too narrowly defined or that the issue or event under study is too recent and, thus, very little research has been written about it. If serious limitations do emerge, consult with your professor about possible ways to overcome them or how to reframe your study.
When discussing the limitations of your research, be sure to:
- Describe each limitation in detailed but concise terms;
- Explain why each limitation exists;
- Provide the reasons why each limitation could not be overcome using the method(s) chosen to gather the data [cite to other studies that had similar problems when possible];
- Assess the impact of each limitation in relation to the overall findings and conclusions of your study; and,
- If appropriate, describe how these limitations could point to the need for further research.
Remember that the method you chose may be the source of a significant limitation that has emerged during your interpretation of the results [for example, you didn't ask a particular question in a survey that you later wish you had]. If this is the case, don't panic. Acknowledge it, and explain how applying a different or more robust methodology might address the research problem more effectively in any future study. A underlying goal of scholarly research is not only to prove what works, but to demonstrate what doesn't work or what needs further clarification.
Brutus, Stéphane et al. Self-Reported Limitations and Future Directions in Scholarly Reports: Analysis and Recommendations. Journal of Management 39 (January 2013): 48-75; Ioannidis, John P.A. Limitations are not Properly Acknowledged in the Scientific Literature. Journal of Clinical Epidemiology 60 (2007): 324-329; Pasek, Josh. Writing the Empirical Social Science Research Paper: A Guide for the Perplexed . January 24, 2012. Academia.edu; Structure: How to Structure the Research Limitations Section of Your Dissertation . Dissertations and Theses: An Online Textbook. Laerd.com; What Is an Academic Paper? Institute for Writing Rhetoric. Dartmouth College; Writing the Experimental Report: Methods, Results, and Discussion. The Writing Lab and The OWL. Purdue University.
Writing Tip
Don't Inflate the Importance of Your Findings! After all the hard work and long hours devoted to writing your research paper, it is easy to get carried away with attributing unwarranted importance to what you’ve done. We all want our academic work to be viewed as excellent and worthy of a good grade, but it is important that you understand and openly acknowledge the limitiations of your study. Inflating of the importance of your study's findings in an attempt hide its flaws is a big turn off to your readers. A measure of humility goes a long way!
Another Writing Tip
Negative Results are Not a Limitation!
Negative evidence refers to findings that unexpectedly challenge rather than support your hypothesis. If you didn't get the results you anticipated, it may mean your hypothesis was incorrect and needs to be reformulated, or, perhaps you have stumbled onto something unexpected that warrants further study. Moreover, the absence of an effect may be very telling in many situations, particularly in experimental research designs. In any case, your results may be of importance to others even though they did not support your hypothesis. Do not fall into the trap of thinking that results contrary to what you expected is a limitation to your study. If you carried out the research well, they are simply your results and only require additional interpretation.
Yet Another Writing Tip
A Note about Sample Size Limitations in Qualitative Research
Sample sizes are typically smaller in qualitative research because, as the study goes on, acquiring more data does not necessarily lead to more information. This is because one occurrence of a piece of data, or a code, is all that is necessary to ensure that it becomes part of the analysis framework. However, it remains true that sample sizes that are too small cannot adequately support claims of having achieved valid conclusions and sample sizes that are too large do not permit the deep, naturalistic, and inductive analysis that defines qualitative inquiry. Determining adequate sample size in qualitative research is ultimately a matter of judgment and experience in evaluating the quality of the information collected against the uses to which it will be applied and the particular research method and purposeful sampling strategy employed. If the sample size is found to be a limitation, it may reflect your judgement about the methodological technique chosen [e.g., single life history study versus focus group interviews] rather than the number of respondents used.
Huberman, A. Michael and Matthew B. Miles. Data Management and Analysis Methods. In Handbook of Qualitative Research. Norman K. Denzin and Yvonna S. Lincoln, eds. (Thousand Oaks, CA: Sage, 1994), pp. 428-444.
- << Previous: 8. The Discussion
- Next: 9. The Conclusion >>
- Last Updated: Jul 18, 2023 11:58 AM
- URL: https://library.sacredheart.edu/c.php?g=29803
- QuickSearch
- Library Catalog
- Databases A-Z
- Publication Finder
- Course Reserves
- Citation Linker
- Digital Commons
- Our Website
Research Support
- Ask a Librarian
- Appointments
- Interlibrary Loan (ILL)
- Research Guides
- Databases by Subject
- Citation Help
Using the Library
- Reserve a Group Study Room
- Renew Books
- Honors Study Rooms
- Off-Campus Access
- Library Policies
- Library Technology
User Information
- Grad Students
- Online Students
- COVID-19 Updates
- Staff Directory
- News & Announcements
- Library Newsletter
My Accounts
- Interlibrary Loan
- Staff Site Login
FIND US ON
How to Write about Research Limitations Without Reducing Your Impact

Being open about what you could not do in your research is actually extremely positive, and it’s viewed favorably by editors and peer reviewers. Writing about your limitations without reducing your impact is a valuable skills that will help your reputation as a researcher.
Areas you might have “failed,” in other words, your limitations, include:
- Aims and objectives (they were a bit too ambitious)
- Study design (not quite right)
- Supporting literature (you’re in uncharted territory)
- Sampling method (if only you’d snowballed it)
- Size of your study population (not enough power)
- Data collection method (bias found its way in)
- Confounding factors (didn’t see that coming!)
Your limitations don’t harm your work and reputation. Quite the opposite, they validate your work and increase your contribution to your field.
Limitations are quite easy to write about in a useful way that won’t reducing your impact. In fact, it’ll increase it.
Why are limitations so important?
Study design limitations, impact limitations, statistical or data limitations, other limitations, how to describe your limitations, where to write your limitations, structure for writing about a limitation, writing up a broader limitation, dealing with breakthroughs and niche-type limitations, dealing with critical flaws, curb your enthusiasm: manage expectations.
Regrettably, the publish-or-perish mentality has created pressure to only come up with successful results. It’s also not too much to say that journals prefer positive studies – where the findings support the hypothesis.
But success alone is not science. Science is trial and error.
So it’s important to present a well-balanced, comprehensive description of your research. That includes your limitations. Accurately reporting your limitations will:
- Help prevent research waste on repeated failures
- Lead to creation of new hypotheses
- Provide useful information for systematic reviews
- Further demonstrate the robustness of your study
Adding clear discussion of any negative results and/or outcomes as well as your study limitations makes you much better able to provide your readers (including peer reviewers ) with:
- Information about your positive results
- Explanation of why your results are credible
- Ideas for future hypothesis generation
- Understanding of why your study has impact
These are good things. There’s even a journal for failure ! That’s how important it is in science.
Some authors find it hard to write about their study limitations, seeing it as an admission of failure. You can do it, and you don’t have to overdo it, either.
Know your limitations and you can anticipate and record them
These might include the procedures, experiments, or reagents (or funding) you have available. As well as specific constraints on the study population. There may be ethical guidelines , and institutional or national policies, that limit what you can do.
These are very common limitations to medical research, for example. We refer to these kinds as study design limitations. Clinical trials, for instance, may have a restriction on interventions expected to have a positive effect. Or there may be restrictions on data collection based on the study population.
Even if your study has a strong design and statistical foundation there might be a strong regional, national, or species-based focus. Or your work could be very population- or experimental-specific.
Your entire field of study, in fact, may only be conducive to incremental findings (e.g., particle physics or molecular biology).
These are inherent limits on impact in that they’re so specific. This limits the extendibility of the findings. It doesn’t however, limit the impact on a specific area or your field. Note the impact and push forward!
Perhaps the most common kind of limitation is statistical or data-based. This category is extremely common in experimental (e.g., chemistry) or field-based (e.g., ecology, population biology, qualitative clinical research) studies.
In many situations, testing hypotheses, you simply may not be able to collect as much data or as good quality data as you want to. Perhaps enrollment was more difficult than expected, under-powering your results.
Statistical limitations can also stem from study design, producing more serious issues in terms of interpreting findings. Seeking expert review from a statistician, such as by using Edanz scientific solutions , may be a good idea before starting your study design.

The above three are often interconnected. And they’re certainly not comprehensive.
As mentioned up top, you may also be limited by the literature. By external confounders. By things you didn’t even see coming (like how long it took you to find 10 qualified respondents for a qualitative study).
Once you’ve identified possible limitations in your work, you need to get to the real point of this post – describing them in your manuscript.
Use the perspective of limitations = contribution and impact to maximize your chances of acceptance.
Reviewers, editors, and readers expect you to present your work authoritatively. You’re the expert in the field, after all. This may make them critical. Embrace that. Counter their possibly negative interpretation by explaining each limitation, showing why the results are still important and useful.
Limitations are usually listed at the end of your Discussion section, though they can also be added throughout. Especially for a long manuscript or for an essay or dissertation, the latter may be useful for the reader.
Writing on your limitations: Words and structure
- This study did have some limitations.
- Three notable limitations affected this study.
- While this study successfully x, there were some limitations.
Giving a specific number is useful for the reader and can guide your writing. But if it’s a longer list, no need to number them. For a short list, you can write them as:
But this gets tiring for more than three limitations (bad RX: reader experience).
So, for longer lists, add a bit of variety in the language to engage the reader. Like this:
- The first issue was…
- Another limitation was…
- Additionally,…
An expert editor will be happy to help you make the English more natural and readable.
After your lead-in sentence, follow a pattern of writing on your findings and related limitation(s), giving a quick interpretation, back it with support (if needed), and offer the next steps.
This provides a complete package for the reader: what happened, what it means, why this is the case, and what is now needed.
In that way, you’ve admitted what may be lacking, but you’ve further established your authority. You’ve also provided a quick roadmap for your reader. That’s an impactful contribution!
It might not always be logical or readable to give that much detail. As long as you fully describe and justify the limitation, you’ve done your job well.
Your study looked at a weight intervention over 6 months at primary healthcare clinics in Japan. The results were generally. But because you only looked at Japanese patients, these findings may not be extendible to patients of other cultures/nationalities, etc.
That’s not a failure at all. It’s a success. But it is a limitation. And other researchers can learn from it and build on it. Write it up in the limitations.
Finding: We found that, in the intervention group, BMI was reduced over 6 months.
Interpretation (and support): This suggests a regimen of routine testing and measurement followed by personalized health guidance from primary physicians had a positive effect on patients’ conditions.
Support: Yamazaki (2019) and Endo et al. (2020) found similar results in urban Japanese clinics and hospitals, respectively.
Limitation and how to use it: While these are useful findings, they are limited by only including Japanese populations. This does not ensure these interventions would be as effective in other nations or cultures. Similar interventions, adapted to the local healthcare and cultural conditions, would help to further clarify the methods.
Now you’ve stated the value of your finding, the limitation, and what to do with it. Nice impact!
Another hurdle you may hit is when your results are particularly novel or you’re publishing in a little-researched field. Those are limitations that need to be stated. In this case, you can support your findings by reinforcing the novelty of your results.
When breaking new ground, there are probably still many gaps in the knowledge base that need to be filled. A good follow-up statement for this type of limitation is to describe what, based on these results, the next steps would be to build a stronger overall evidence base.

It’s possible that your study will have a fairly “critical” flaw (usually in the study design) that decreases confidence in your findings.
Other experts will likely notice this (in peer review or perhaps on a preprint server, they should notice it), so it’s best to explain why this error or flaw occurred.
You can still explain why the study is worth repeating or how you plan to retest the phenomenon. But you may need to temper your publication goals if you still plan to publish your work.
No one expects science to be perfect the first time and while your peers can be highly critical, no one’s work is beyond limitations. This is important to keep in mind.
Edanz experts can help by giving you an Expert Scientific Review and seeking out your limitations.
Our knowledge base is built on uncovering each piece of the puzzle, one at a time, and limitations show us where new efforts need to be made. Much like peer review , don’t think of limitations as being inherently bad, but more as an opportunity for a new challenge.
Ultimately, your limitations may be someone else’s inspirations. Include them in your submission when you get published in the journal of your choice .
All research faces problems: Being honest impresses people much more than ignoring your limitations.

- Researcher Services
- English Editing
- EXCITED by the SCIENCE
- Smart Tools
- Journal Selector
- About Edanz
- Privacy Policy
- Terms & Conditions
- Services & Pricing
- 特定商取引法に基づく表記

CPS Online Graduate Studies Research Paper (UNH Manchester Library): Limitations of the Study
- Overview of the Research Process for Capstone Projects
- Types of Research Design
- Selecting a Research Problem
- The Title of Your Research Paper
- Before You Begin Writing
- 7 Parts of the Research Paper
- Background Information
- Quanitative and Qualitative Methods
- Qualitative Methods
- Quanitative Methods
- Resources to Help You With the Literature Review
- Non-Textual Elements
Limitations of the Study
- Format of Capstone Research Projects at GSC
- Editing and Proofreading Your Paper
- Acknowledgements
- UNH Scholar's Repository
The limitations of the study are those characteristics of design or methodology that impacted or influenced the interpretation of the findings from your research. They are the constraints on generalizability, applications to practice, and/or utility of findings that are the result of the ways in which you initially chose to design the study and/or the method used to establish internal and external validity.
Price, James H. and Judy Murnan. “Research Limitations and the Necessity of Reporting Them.” American Journal of Health Education 35 (2004): 66-67.
Always acknowledge a study's limitations. It is far better that you identify and acknowledge your study’s limitations than to have them pointed out by your professor and be graded down because you appear to have ignored them.
Keep in mind that acknowledgement of a study's limitations is an opportunity to make suggestions for further research. If you do connect your study's limitations to suggestions for further research, be sure to explain the ways in which these unanswered questions may become more focused because of your study.
Acknowledgement of a study's limitations also provides you with an opportunity to demonstrate that you have thought critically about the research problem, understood the relevant literature published about it, and correctly assessed the methods chosen for studying the problem. A key objective of the research process is not only discovering new knowledge but to also confront assumptions and explore what we don't know.
Claiming limitations is a subjective process because you must evaluate the impact of those limitations . Don't just list key weaknesses and the magnitude of a study's limitations. To do so diminishes the validity of your research because it leaves the reader wondering whether, or in what ways, limitation(s) in your study may have impacted the results and conclusions. Limitations require a critical, overall appraisal and interpretation of their impact. You should answer the question: do these problems with errors, methods, validity, etc. eventually matter and, if so, to what extent?
Price, James H. and Judy Murnan. “Research Limitations and the Necessity of Reporting Them.” American Journal of Health Education 35 (2004): 66-67; Structure: How to Structure the Research Limitations Section of Your Dissertation . Dissertations and Theses: An Online Textbook. Laerd.com.
Descriptions of Possible Limitations
All studies have limitations . However, it is important that you restrict your discussion to limitations related to the research problem under investigation. For example, if a meta-analysis of existing literature is not a stated purpose of your research, it should not be discussed as a limitation. Do not apologize for not addressing issues that you did not promise to investigate in the introduction of your paper.
Here are examples of limitations related to methodology and the research process you may need to describe and to discuss how they possibly impacted your results. Descriptions of limitations should be stated in the past tense because they were discovered after you completed your research.
Possible Methodological Limitations
- Sample size -- the number of the units of analysis you use in your study is dictated by the type of research problem you are investigating. Note that, if your sample size is too small, it will be difficult to find significant relationships from the data, as statistical tests normally require a larger sample size to ensure a representative distribution of the population and to be considered representative of groups of people to whom results will be generalized or transferred. Note that sample size is less relevant in qualitative research.
- Lack of available and/or reliable data -- a lack of data or of reliable data will likely require you to limit the scope of your analysis, the size of your sample, or it can be a significant obstacle in finding a trend and a meaningful relationship. You need to not only describe these limitations but to offer reasons why you believe data is missing or is unreliable. However, don’t just throw up your hands in frustration; use this as an opportunity to describe the need for future research.
- Lack of prior research studies on the topic -- citing prior research studies forms the basis of your literature review and helps lay a foundation for understanding the research problem you are investigating. Depending on the currency or scope of your research topic, there may be little, if any, prior research on your topic. Before assuming this to be true, though, consult with a librarian. In cases when a librarian has confirmed that there is no prior research, you may be required to develop an entirely new research typology [for example, using an exploratory rather than an explanatory research design]. Note again that discovering a limitation can serve as an important opportunity to identify new gaps in the literature and to describe the need for further research.
- Measure used to collect the data -- sometimes it is the case that, after completing your interpretation of the findings, you discover that the way in which you gathered data inhibited your ability to conduct a thorough analysis of the results. For example, you regret not including a specific question in a survey that, in retrospect, could have helped address a particular issue that emerged later in the study. Acknowledge the deficiency by stating a need for future researchers to revise the specific method for gathering data.
- Self-reported data -- whether you are relying on pre-existing data or you are conducting a qualitative research study and gathering the data yourself, self-reported data is limited by the fact that it rarely can be independently verified. In other words, you have to take what people say, whether in interviews, focus groups, or on questionnaires, at face value. However, self-reported data can contain several potential sources of bias that you should be alert to and note as limitations. These biases become apparent if they are incongruent with data from other sources. These are: (1) selective memory [remembering or not remembering experiences or events that occurred at some point in the past]; (2) telescoping [recalling events that occurred at one time as if they occurred at another time]; (3) attribution [the act of attributing positive events and outcomes to one's own agency but attributing negative events and outcomes to external forces]; and, (4) exaggeration [the act of representing outcomes or embellishing events as more significant than is actually suggested from other data].
Possible Limitations of the Researcher
- Access -- if your study depends on having access to people, organizations, or documents and, for whatever reason, access is denied or limited in some way, the reasons for this need to be described.
- Longitudinal effects -- unlike your professor, who can literally devote years [even a lifetime] to studying a single topic, the time available to investigate a research problem and to measure change or stability over time is pretty much constrained by the due date of your assignment. Be sure to choose a research problem that does not require an excessive amount of time to complete the literature review, apply the methodology, and gather and interpret the results. If you're unsure whether you can complete your research within the confines of the assignment's due date, talk to your professor.
- Cultural and other type of bias -- we all have biases, whether we are conscience of them or not. Bias is when a person, place, or thing is viewed or shown in a consistently inaccurate way. Bias is usually negative, though one can have a positive bias as well, especially if that bias reflects your reliance on research that only support for your hypothesis. When proof-reading your paper, be especially critical in reviewing how you have stated a problem, selected the data to be studied, what may have been omitted, the manner in which you have ordered events, people, or places, how you have chosen to represent a person, place, or thing, to name a phenomenon, or to use possible words with a positive or negative connotation.
NOTE: If you detect bias in prior research, it must be acknowledged and you should explain what measures were taken to avoid perpetuating that bias.
- Fluency in a language -- if your research focuses on measuring the perceived value of after-school tutoring among Mexican-American ESL [English as a Second Language] students, for example, and you are not fluent in Spanish, you are limited in being able to read and interpret Spanish language research studies on the topic. This deficiency should be acknowledged.
Aguinis, Hermam and Jeffrey R. Edwards. “Methodological Wishes for the Next Decade and How to Make Wishes Come True.” Journal of Management Studies 51 (January 2014): 143-174; Brutus, Stéphane et al. "Self-Reported Limitations and Future Directions in Scholarly Reports: Analysis and Recommendations." Journal of Management 39 (January 2013): 48-75; Senunyeme, Emmanuel K. Business Research Methods . Powerpoint Presentation. Regent University of Science and Technology; ter Riet, Gerben et al. “All That Glitters Isn't Gold: A Survey on Acknowledgment of Limitations in Biomedical Studies.” PLOS One 8 (November 2013): 1-6.
Structure and Writing Style
Information about the limitations of your study are generally placed either at the beginning of the discussion section of your paper so the reader knows and understands the limitations before reading the rest of your analysis of the findings, or, the limitations are outlined at the conclusion of the discussion section as an acknowledgement of the need for further study. Statements about a study's limitations should not be buried in the body [middle] of the discussion section unless a limitation is specific to something covered in that part of the paper. If this is the case, though, the limitation should be reiterated at the conclusion of the section. If you determine that your study is seriously flawed due to important limitations, such as, an inability to acquire critical data, consider reframing it as an exploratory study intended to lay the groundwork for a more complete research study in the future. Be sure, though, to specifically explain the ways that these flaws can be successfully overcome in a new study. But, do not use this as an excuse for not developing a thorough research paper! Review the tab in this guide for developing a research topic. If serious limitations exist, it generally indicates a likelihood that your research problem is too narrowly defined or that the issue or event under study is too recent and, thus, very little research has been written about it. If serious limitations do emerge, consult with your professor about possible ways to overcome them or how to revise your study. When discussing the limitations of your research, be sure to: Describe each limitation in detailed but concise terms; Explain why each limitation exists; Provide the reasons why each limitation could not be overcome using the method(s) chosen to acquire or gather the data [cite to other studies that had similar problems when possible]; Assess the impact of each limitation in relation to the overall findings and conclusions of your study; and, If appropriate, describe how these limitations could point to the need for further research. Remember that the method you chose may be the source of a significant limitation that has emerged during your interpretation of the results [for example, you didn't interview a group of people that you later wish you had]. If this is the case, don't panic. Acknowledge it, and explain how applying a different or more robust methodology might address the research problem more effectively in a future study. A underlying goal of scholarly research is not only to show what works, but to demonstrate what doesn't work or what needs further clarification. Aguinis, Hermam and Jeffrey R. Edwards. “Methodological Wishes for the Next Decade and How to Make Wishes Come True.” Journal of Management Studies 51 (January 2014): 143-174; Brutus, Stéphane et al. "Self-Reported Limitations and Future Directions in Scholarly Reports: Analysis and Recommendations." Journal of Management 39 (January 2013): 48-75; Ioannidis, John P.A. "Limitations are not Properly Acknowledged in the Scientific Literature." Journal of Clinical Epidemiology 60 (2007): 324-329; Pasek, Josh. Writing the Empirical Social Science Research Paper: A Guide for the Perplexed. January 24, 2012. Academia.edu; Structure: How to Structure the Research Limitations Section of Your Dissertation. Dissertations and Theses: An Online Textbook. Laerd.com; What Is an Academic Paper? Institute for Writing Rhetoric. Dartmouth College; Writing the Experimental Report: Methods, Results, and Discussion. The Writing Lab and The OWL. Purdue University.
Information about the limitations of your study are generally placed either at the beginning of the discussion section of your paper so the reader knows and understands the limitations before reading the rest of your analysis of the findings, or, the limitations are outlined at the conclusion of the discussion section as an acknowledgement of the need for further study. Statements about a study's limitations should not be buried in the body [middle] of the discussion section unless a limitation is specific to something covered in that part of the paper. If this is the case, though, the limitation should be reiterated at the conclusion of the section.
If you determine that your study is seriously flawed due to important limitations , such as, an inability to acquire critical data, consider reframing it as an exploratory study intended to lay the groundwork for a more complete research study in the future. Be sure, though, to specifically explain the ways that these flaws can be successfully overcome in a new study.
But, do not use this as an excuse for not developing a thorough research paper! Review the tab in this guide for developing a research topic . If serious limitations exist, it generally indicates a likelihood that your research problem is too narrowly defined or that the issue or event under study is too recent and, thus, very little research has been written about it. If serious limitations do emerge, consult with your professor about possible ways to overcome them or how to revise your study.
When discussing the limitations of your research, be sure to:
- Describe each limitation in detailed but concise terms;
- Explain why each limitation exists;
- Provide the reasons why each limitation could not be overcome using the method(s) chosen to acquire or gather the data [cite to other studies that had similar problems when possible];
- Assess the impact of each limitation in relation to the overall findings and conclusions of your study; and,
- If appropriate, describe how these limitations could point to the need for further research.
Remember that the method you chose may be the source of a significant limitation that has emerged during your interpretation of the results [for example, you didn't interview a group of people that you later wish you had]. If this is the case, don't panic. Acknowledge it, and explain how applying a different or more robust methodology might address the research problem more effectively in a future study. A underlying goal of scholarly research is not only to show what works, but to demonstrate what doesn't work or what needs further clarification.
Aguinis, Hermam and Jeffrey R. Edwards. “Methodological Wishes for the Next Decade and How to Make Wishes Come True.” Journal of Management Studies 51 (January 2014): 143-174; Brutus, Stéphane et al. "Self-Reported Limitations and Future Directions in Scholarly Reports: Analysis and Recommendations." Journal of Management 39 (January 2013): 48-75; Ioannidis, John P.A. "Limitations are not Properly Acknowledged in the Scientific Literature." Journal of Clinical Epidemiology 60 (2007): 324-329; Pasek, Josh. Writing the Empirical Social Science Research Paper: A Guide for the Perplexed . January 24, 2012. Academia.edu; Structure: How to Structure the Research Limitations Section of Your Dissertation . Dissertations and Theses: An Online Textbook. Laerd.com; What Is an Academic Paper? Institute for Writing Rhetoric. Dartmouth College; Writing the Experimental Report: Methods, Results, and Discussion . The Writing Lab and The OWL. Purdue University.
- << Previous: The Discussion
- Next: Conclusion >>
- Last Updated: Nov 6, 2023 1:43 PM
- URL: https://libraryguides.unh.edu/cpsonlinegradpaper
Research Limitations & Delimitations
What they are and how they’re different (with examples)
By: Derek Jansen (MBA) | Expert Reviewed By: David Phair (PhD) | September 2022
If you’re new to the world of research, you’ve probably heard the terms “ research limitations ” and “ research delimitations ” being thrown around, often quite loosely. In this post, we’ll unpack what both of these mean, how they’re similar and how they’re different – so that you can write up these sections the right way.
Overview: Limitations vs Delimitations
- Are they the same?
- What are research limitations
- What are research delimitations
- Limitations vs delimitations
First things first…
Let’s start with the most important takeaway point of this post – research limitations and research delimitations are not the same – but they are related to each other (we’ll unpack that a little later). So, if you hear someone using these two words interchangeably, be sure to share this post with them!
Research Limitations
Research limitations are, at the simplest level, the weaknesses of the study , based on factors that are often outside of your control as the researcher. These factors could include things like time , access to funding, equipment , data or participants . For example, if you weren’t able to access a random sample of participants for your study and had to adopt a convenience sampling strategy instead, that would impact the generalizability of your findings and therefore reflect a limitation of your study.
Research limitations can also emerge from the research design itself . For example, if you were undertaking a correlational study, you wouldn’t be able to infer causality (since correlation doesn’t mean certain causation). Similarly, if you utilised online surveys to collect data from your participants, you naturally wouldn’t be able to get the same degree of rich data that you would from in-person interviews .
Simply put, research limitations reflect the shortcomings of a study , based on practical (or theoretical) constraints that the researcher faced. These shortcomings limit what you can conclude from a study, but at the same time, present a foundation for future research . Importantly, all research has limitations , so there’s no need to hide anything here – as long as you discuss how the limitations might affect your findings, it’s all good.
Research Delimitations
Alright, now that we’ve unpacked the limitations, let’s move on to the delimitations .
Research delimitations are similar to limitations in that they also “ limit ” the study, but their focus is entirely different. Specifically, the delimitations of a study refer to the scope of the research aims and research questions . In other words, delimitations reflect the choices you, as the researcher, intentionally make in terms of what you will and won’t try to achieve with your study. In other words, what your research aims and research questions will and won’t include.
As we’ve spoken about many times before, it’s important to have a tight, narrow focus for your research, so that you can dive deeply into your topic, apply your energy to one specific area and develop meaningful insights. If you have an overly broad scope or unfocused topic, your research will often pull in multiple, even opposing directions, and you’ll just land up with a muddy mess of findings .
So, the delimitations section is where you’ll clearly state what your research aims and research questions will focus on – and just as importantly, what they will exclude . For example, you might investigate a widespread phenomenon, but choose to focus your study on a specific age group, ethnicity or gender. Similarly, your study may focus exclusively on one country, city or even organization. As long as the scope is well justified (in other words, it represents a novel, valuable research topic), this is perfectly acceptable – in fact, it’s essential. Remember, focus is your friend.
Need a helping hand?
Conclusion: Limitations vs Delimitations
Ok, so let’s recap.
Research limitations and research delimitations are related in that they both refer to “limits” within a study. But, they are distinctly different. Limitations reflect the shortcomings of your study, based on practical or theoretical constraints that you faced.
Contrasted to that, delimitations reflect the choices that you made in terms of the focus and scope of your research aims and research questions. If you want to learn more about research aims and questions, you can check out this video post , where we unpack those concepts in detail.

Psst... there’s more!
This post was based on one of our popular Research Bootcamps . If you're working on a research project, you'll definitely want to check this out ...
You Might Also Like:

18 Comments
Good clarification of ideas on how a researcher ought to do during Process of choice
Thank you so much for this very simple but explicit explanation on limitation and delimitation. It has so helped me to develop my masters proposal. hope to recieve more from your site as time progresses
Thank you for this explanation – very clear.
Thanks for the explanation, really got it well.
This website is really helpful for my masters proposal
Thank you very much for helping to explain these two terms
I spent almost the whole day trying to figure out the differences
when I came across your notes everything became very clear
thanks for the clearly outlined explanation on the two terms, limitation and delimitation.
Very helpful Many thanks 🙏
Excellent it resolved my conflict .
I would like you to assist me please. If in my Research, I interviewed some participants and I submitted Questionnaires to other participants to answered to the questions, in the same organization, Is this a Qualitative methodology , a Quantitative Methodology or is it a Mixture Methodology I have used in my research? Please help me
How do I cite this article in APA format
Really so great ,finally have understood it’s difference now
Getting more clear regarding Limitations and Delimitation and concepts
I really appreciate your apt and precise explanation of the two concepts namely ; Limitations and Delimitations.
This is a good sources of research information for learners.
thank you for this, very helpful to researchers
Very good explained
Great and clear explanation, after a long confusion period on the two words, i can now explain to someone with ease.
Submit a Comment Cancel reply
Your email address will not be published. Required fields are marked *
Save my name, email, and website in this browser for the next time I comment.
- Print Friendly
- Link to facebook
- Link to linkedin
- Link to twitter
- Link to youtube
- Writing Tips
How to Identify Limitations in Research

4-minute read
- 7th March 2022
Whether you’re a veteran researcher with years of experience under your belt or a novice to the field that’s feeling overwhelmed with where to start, you must understand that every study has its limitations. These are restrictions that arise from the study’s design, or the methodology implemented during the testing phase. Unfortunately, research limitations will always exist due to the subjective nature of testing a hypothesis. We’ve compiled some helpful information below on how to identify and accept research limitations and use them to your advantage. Essentially, we’ll show you how to make lemonade (a brilliant piece of academic work ) from the lemons you receive (the constraints your study reveals).
Research Limitations
So, let’s dive straight in, shall we? It’s always beneficial (and good practice) to disclose your research limitations . A common thought is that divulging these shortcomings will undermine the credibility and quality of your research. However, this is certainly not the case— stating the facts upfront not only reinforces your reputation as a researcher but also lets the assessor or reader know that you’re confident and transparent about the results and relevance of your study, despite these constraints.
Additionally, it creates a gap for more research opportunities, where you can analyze these limitations and determine how to incorporate or address them in a new batch of tests or create a new hypothesis altogether. Another bonus is that it helps readers to understand the optimum conditions for how to apply the results of your testing. This is a win-win, making for a far more persuasive research paper .
Now that you know why you should clarify your research limitations, let’s focus on which ones take precedence and should be disclosed. Any given research project can be vulnerable to various hindrances, so how do you identify them and single out the most significant ones to discuss? Well, that depends entirely on the nature of your study. You’ll need to comb through your research approach, methodology, testing processes, and expected results to identify the type of limitations your study may be exposed to. It’s worth noting that this understanding can only offer a broad idea of the possible restrictions you’ll face and may potentially change throughout the study.
We’ve compiled a list of the most common types of research limitations that you may encounter so you can adequately prepare for them and remain vigilant during each stage of your study.
Sample Size:
It’s critical that you choose a sample size that accurately represents the population you wish to test your theory on. If a sample is too small, the results cannot reliably be generalized across a large population.
Methodology:
The method you choose before you commence testing might seem effective in theory, but too many stumbling blocks during the testing phase can influence the accuracy and reliability of the results.
Find this useful?
Subscribe to our newsletter and get writing tips from our editors straight to your inbox.
Collection of Data:
The methods you utilize to obtain your research—surveys, emails, in-person interviews, phone calls—will directly influence the type of results your study yields.
Age of Data:
The nature of the information—and how far back it goes—affects the type of assumptions you can make. Extrapolating older data for a current hypothesis can significantly change the outcome of your testing.
Time Constraints:
Working within the deadline of when you need to submit your findings will determine the extent of your research and testing and, therefore, can heavily impact your results. Limited time frames for testing might mean not achieving the scope of results you were originally looking for.
Limited Budget:
Your study may require equipment and other resources that can become extremely costly. Budget constraints may mean you cannot acquire advanced software, programs, or travel to multiple destinations to interview participants. All of these factors can substantially influence your results.
So, now that you know how to determine your research limitations and the types you might experience, where should you document it? It’s commonly disclosed at the beginning of your discussion section , so the reader understands the shortcomings of your study before digging into the juicy bit—your findings. Alternatively, you can detail the constraints faced at the end of the discussion section to emphasize the requirements for the completion of further studies.
We hope this post will prepare you for some of the pitfalls you may encounter when conducting and documenting your research. Once you have a first draft ready, consider submitting a free sample to us for proofreading to ensure that your writing is concise and error-free and your results—despite their limitations— shine through.
Share this article:
Post A New Comment
Got content that needs a quick turnaround? Let us polish your work. Explore our editorial business services.
9-minute read
How to Use Infographics to Boost Your Presentation
Is your content getting noticed? Capturing and maintaining an audience’s attention is a challenge when...
8-minute read
Why Interactive PDFs Are Better for Engagement
Are you looking to enhance engagement and captivate your audience through your professional documents? Interactive...
7-minute read
Seven Key Strategies for Voice Search Optimization
Voice search optimization is rapidly shaping the digital landscape, requiring content professionals to adapt their...
Five Creative Ways to Showcase Your Digital Portfolio
Are you a creative freelancer looking to make a lasting impression on potential clients or...
How to Ace Slack Messaging for Contractors and Freelancers
Effective professional communication is an important skill for contractors and freelancers navigating remote work environments....
3-minute read
How to Insert a Text Box in a Google Doc
Google Docs is a powerful collaborative tool, and mastering its features can significantly enhance your...

Make sure your writing is the best it can be with our expert English proofreading and editing.

Research Limitations
It is for sure that your research will have some limitations and it is normal. However, it is critically important for you to be striving to minimize the range of scope of limitations throughout the research process. Also, you need to provide the acknowledgement of your research limitations in conclusions chapter honestly.
It is always better to identify and acknowledge shortcomings of your work, rather than to leave them pointed out to your by your dissertation assessor. While discussing your research limitations, don’t just provide the list and description of shortcomings of your work. It is also important for you to explain how these limitations have impacted your research findings.
Your research may have multiple limitations, but you need to discuss only those limitations that directly relate to your research problems. For example, if conducting a meta-analysis of the secondary data has not been stated as your research objective, no need to mention it as your research limitation.
Research limitations in a typical dissertation may relate to the following points:
1. Formulation of research aims and objectives . You might have formulated research aims and objectives too broadly. You can specify in which ways the formulation of research aims and objectives could be narrowed so that the level of focus of the study could be increased.
2. Implementation of data collection method . Because you do not have an extensive experience in primary data collection (otherwise you would not be reading this book), there is a great chance that the nature of implementation of data collection method is flawed.
3. Sample size. Sample size depends on the nature of the research problem. If sample size is too small, statistical tests would not be able to identify significant relationships within data set. You can state that basing your study in larger sample size could have generated more accurate results. The importance of sample size is greater in quantitative studies compared to qualitative studies.
4. Lack of previous studies in the research area . Literature review is an important part of any research, because it helps to identify the scope of works that have been done so far in research area. Literature review findings are used as the foundation for the researcher to be built upon to achieve her research objectives.
However, there may be little, if any, prior research on your topic if you have focused on the most contemporary and evolving research problem or too narrow research problem. For example, if you have chosen to explore the role of Bitcoins as the future currency, you may not be able to find tons of scholarly paper addressing the research problem, because Bitcoins are only a recent phenomenon.
5. Scope of discussions . You can include this point as a limitation of your research regardless of the choice of the research area. Because (most likely) you don’t have many years of experience of conducing researches and producing academic papers of such a large size individually, the scope and depth of discussions in your paper is compromised in many levels compared to the works of experienced scholars.
You can discuss certain points from your research limitations as the suggestion for further research at conclusions chapter of your dissertation.
My e-book, The Ultimate Guide to Writing a Dissertation in Business Studies: a step by step assistance offers practical assistance to complete a dissertation with minimum or no stress. The e-book covers all stages of writing a dissertation starting from the selection to the research area to submitting the completed version of the work within the deadline. John Dudovskiy

- Cookies & Privacy
- GETTING STARTED
- Introduction
- FUNDAMENTALS
- Acknowledgements
- Research questions & hypotheses
- Concepts, constructs & variables
- Research limitations
- Getting started
- Sampling Strategy
- Research Quality
- Research Ethics
- Data Analysis
How to structure the Research Limitations section of your dissertation
There is no "one best way" to structure the Research Limitations section of your dissertation. However, we recommend a structure based on three moves : the announcing , reflecting and forward looking move. The announcing move immediately allows you to identify the limitations of your dissertation and explain how important each of these limitations is. The reflecting move provides greater depth, helping to explain the nature of the limitations and justify the choices that you made during the research process. Finally, the forward looking move enables you to suggest how such limitations could be overcome in future. The collective aim of these three moves is to help you walk the reader through your Research Limitations section in a succinct and structured way. This will make it clear to the reader that you recognise the limitations of your own research, that you understand why such factors are limitations, and can point to ways of combating these limitations if future research was carried out. This article explains what should be included in each of these three moves :
- THE ANNOUNCING MOVE: Identifying limitations and explaining how important they are
- THE REFLECTING MOVE: Explaining the nature of the limitations and justifying the choices you made
- THE FORWARD LOOKING MOVE: Suggesting how such limitations could be overcome in future
THE ANNOUNCING MOVE Identifying limitations, and explaining how important they are
There are many possible limitations that your research may have faced. However, is not necessary for you to discuss all of these limitations in your Research Limitations section. After all, you are not writing a 2000 word critical review of the limitations of your dissertation, just a 200-500 word critique that is only one section long (i.e., the Research Limitations section within your Conclusions chapter). Therefore, in this first announcing move , we would recommend that you identify only those limitations that had the greatest potential impact on: (a) the quality of your findings; and (b) your ability to effectively answer your research questions and/or hypotheses.
We use the word potential impact because we often do not know the degree to which different factors limited our findings or our ability to effectively answer our research questions and/or hypotheses. For example, we know that when adopting a quantitative research design, a failure to use a probability sampling technique significantly limits our ability to make broader generalisations from our results (i.e., our ability to make statistical inferences from our sample to the population being studied). However, the degree to which this reduces the quality of our findings is a matter of debate. Also, whilst the lack of a probability sampling technique when using a quantitative research design is a very obvious example of a research limitation, other limitations are far less clear. Therefore, the key point is to focus on those limitations that you feel had the greatest impact on your findings, as well as your ability to effectively answer your research questions and/or hypotheses.
Overall, the announcing move should be around 10-20% of the total word count of the Research Limitations section.
THE REFLECTING MOVE Explaining the nature of the limitations and justifying the choices you made
Having identified the most important limitations to your dissertation in the announcing move , the reflecting move focuses on explaining the nature of these limitations and justifying the choices that you made during the research process. This part should be around 60-70% of the total word count of the Research Limitations section.
It is important to remember at this stage that all research suffers from limitations, whether it is performed by undergraduate and master's level dissertation students, or seasoned academics. Acknowledging such limitations should not be viewed as a weakness, highlighting to the person marking your work the reasons why you should receive a lower grade. Instead, the reader is more likely to accept that you recognise the limitations of your own research if you write a high quality reflecting move . This is because explaining the limitations of your research and justifying the choices you made during the dissertation process demonstrates the command that you had over your research.
We talk about explaining the nature of the limitations in your dissertation because such limitations are highly research specific. Let's take the example of potential limitations to your sampling strategy. Whilst you may have a number of potential limitations in sampling strategy, let's focus on the lack of probability sampling ; that is, of all the different types of sampling technique that you could have used [see Types of probability sampling and Types of non-probability sampling ], you choose not to use a probability sampling technique (e.g., simple random sampling , systematic random sampling , stratified random sampling ). As mentioned, if you used a quantitative research design in your dissertation, the lack of probability sampling is an important, obvious limitation to your research. This is because it prevents you from making generalisations about the population you are studying (e.g. Facebook usage at a single university of 20,000 students) from the data you have collected (e.g., a survey of 400 students at the same university). Since an important component of quantitative research is such generalisation, this is a clear limitation. However, the lack of a probability sampling technique is not viewed as a limitation if you used a qualitative research design. In qualitative research designs, a non-probability sampling technique is typically selected over a probability sampling technique.
And this is just part of the puzzle?
Even if you used a quantitative research design, but failed to employ a probability sampling technique, there are still many perfectly justifiable reasons why you could have made such a choice. For example, it may have been impossible (or near on impossible) to get a list of the population you were studying (e.g., a list of all the 20,000 students at the single university you were interested in). Since probability sampling is only possible when we have such a list, the lack of such a list or inability to attain such a list is a perfectly justifiable reason for not using a probability sampling technique; even if such a technique is the ideal.
As such, the purpose of all the guides we have written on research limitations is to help you: (a) explain the nature of the limitations in your dissertation; and (b) justify the choices you made.
In helping you to justifying the choices that you made, these articles explain not only when something is, in theory , an obvious limitation, but how, in practice , such a limitation was not necessarily so damaging to the quality of your dissertation. This should significantly strengthen the quality of your Research Limitations section.
THE FORWARD LOOKING MOVE Suggesting how such limitations could be overcome in future
Finally, the forward looking move builds on the reflecting move by suggesting how the limitations you have discuss could be overcome through future research. Whilst a lot could be written in this part of the Research Limitations section, we would recommend that it is only around 10-20% of the total word count for this section.
Educational resources and simple solutions for your research journey

How to Present the Limitations of a Study in Research?
The limitations of the study convey to the reader how and under which conditions your study results will be evaluated. Scientific research involves investigating research topics, both known and unknown, which inherently includes an element of risk. The risk could arise due to human errors, barriers to data gathering, limited availability of resources, and researcher bias. Researchers are encouraged to discuss the limitations of their research to enhance the process of research, as well as to allow readers to gain an understanding of the study’s framework and value.
Limitations of the research are the constraints placed on the ability to generalize from the results and to further describe applications to practice. It is related to the utility value of the findings based on how you initially chose to design the study, the method used to establish internal and external validity, or the result of unanticipated challenges that emerged during the study. Knowing about these limitations and their impact can explain how the limitations of your study can affect the conclusions and thoughts drawn from your research. 1
Table of Contents
What are the limitations of a study
Researchers are probably cautious to acknowledge what the limitations of the research can be for fear of undermining the validity of the research findings. No research can be faultless or cover all possible conditions. These limitations of your research appear probably due to constraints on methodology or research design and influence the interpretation of your research’s ultimate findings. 2 These are limitations on the generalization and usability of findings that emerge from the design of the research and/or the method employed to ensure validity internally and externally. But such limitations of the study can impact the whole study or research paper. However, most researchers prefer not to discuss the different types of limitations in research for fear of decreasing the value of their paper amongst the reviewers or readers.

Importance of limitations of a study
Writing the limitations of the research papers is often assumed to require lots of effort. However, identifying the limitations of the study can help structure the research better. Therefore, do not underestimate the importance of research study limitations. 3
- Opportunity to make suggestions for further research. Suggestions for future research and avenues for further exploration can be developed based on the limitations of the study.
- Opportunity to demonstrate critical thinking. A key objective of the research process is to discover new knowledge while questioning existing assumptions and exploring what is new in the particular field. Describing the limitation of the research shows that you have critically thought about the research problem, reviewed relevant literature, and correctly assessed the methods chosen for studying the problem.
- Demonstrate Subjective learning process. Writing limitations of the research helps to critically evaluate the impact of the said limitations, assess the strength of the research, and consider alternative explanations or interpretations. Subjective evaluation contributes to a more complex and comprehensive knowledge of the issue under study.
Why should I include limitations of research in my paper
All studies have limitations to some extent. Including limitations of the study in your paper demonstrates the researchers’ comprehensive and holistic understanding of the research process and topic. The major advantages are the following:
- Understand the study conditions and challenges encountered . It establishes a complete and potentially logical depiction of the research. The boundaries of the study can be established, and realistic expectations for the findings can be set. They can also help to clarify what the study is not intended to address.
- Improve the quality and validity of the research findings. Mentioning limitations of the research creates opportunities for the original author and other researchers to undertake future studies to improve the research outcomes.
- Transparency and accountability. Including limitations of the research helps maintain mutual integrity and promote further progress in similar studies.
- Identify potential bias sources. Identifying the limitations of the study can help researchers identify potential sources of bias in their research design, data collection, or analysis. This can help to improve the validity and reliability of the findings.
Where do I need to add the limitations of the study in my paper
The limitations of your research can be stated at the beginning of the discussion section, which allows the reader to comprehend the limitations of the study prior to reading the rest of your findings or at the end of the discussion section as an acknowledgment of the need for further research.
Types of limitations in research
There are different types of limitations in research that researchers may encounter. These are listed below:
- Research Design Limitations : Restrictions on your research or available procedures may affect the research outputs. If the research goals and objectives are too broad, explain how they should be narrowed down to enhance the focus of your study. If there was a selection bias in your sample, explain how this may affect the generalizability of your findings. This can help readers understand the limitations of the study in terms of their impact on the overall validity of your research.
- Impact Limitations : Your study might be limited by a strong regional-, national-, or species-based impact or population- or experimental-specific impact. These inherent limitations on impact affect the extendibility and generalizability of the findings.
- Data or statistical limitations : Data or statistical limitations in research are extremely common in experimental (such as medicine, physics, and chemistry) or field-based (such as ecology and qualitative clinical research) studies. Sometimes, it is either extremely difficult to acquire sufficient data or gain access to the data. These limitations of the research might also be the result of your study’s design and might result in an incomplete conclusion to your research.
Limitations of study examples
All possible limitations of the study cannot be included in the discussion section of the research paper or dissertation. It will vary greatly depending on the type and nature of the study. These include types of research limitations that are related to methodology and the research process and that of the researcher as well that you need to describe and discuss how they possibly impacted your results.
Common methodological limitations of the study
Limitations of research due to methodological problems are addressed by identifying the potential problem and suggesting ways in which this should have been addressed. Some potential methodological limitations of the study are as follows. 1
- Sample size: The sample size 4 is dictated by the type of research problem investigated. If the sample size is too small, finding a significant relationship from the data will be difficult, as statistical tests require a large sample size to ensure a representative population distribution and generalize the study findings.
- Lack of available/reliable data: A lack of available/reliable data will limit the scope of your analysis and the size of your sample or present obstacles in finding a trend or meaningful relationship. So, when writing about the limitations of the study, give convincing reasons why you feel data is absent or untrustworthy and highlight the necessity for a future study focused on developing a new data-gathering strategy.
- Lack of prior research studies: Citing prior research studies is required to help understand the research problem being investigated. If there is little or no prior research, an exploratory rather than an explanatory research design will be required. Also, discovering the limitations of the study presents an opportunity to identify gaps in the literature and describe the need for additional study.
- Measure used to collect the data: Sometimes, the data gathered will be insufficient to conduct a thorough analysis of the results. A limitation of the study example, for instance, is identifying in retrospect that a specific question could have helped address a particular issue that emerged during data analysis. You can acknowledge the limitation of the research by stating the need to revise the specific method for gathering data in the future.
- Self-reported data: Self-reported data cannot be independently verified and can contain several potential bias sources, such as selective memory, attribution, and exaggeration. These biases become apparent if they are incongruent with data from other sources.
General limitations of researchers
Limitations related to the researcher can also influence the study outcomes. These should be addressed, and related remedies should be proposed.
- Limited access to data : If your study requires access to people, organizations, data, or documents whose access is denied or limited, the reasons need to be described. An additional explanation stating why this limitation of research did not prevent you from following through on your study is also needed.
- Time constraints : Researchers might also face challenges in meeting research deadlines due to a lack of timely participant availability or funds, among others. The impacts of time constraints must be acknowledged by mentioning the need for a future study addressing this research problem.
- Conflicts due to biased views and personal issues : Differences in culture or personal views can contribute to researcher bias, as they focus only on the results and data that support their main arguments. To avoid this, pay attention to the problem statement and data gathering.
Steps for structuring the limitations section
Limitations are an inherent part of any research study. Issues may vary, ranging from sampling and literature review to methodology and bias. However, there is a structure for identifying these elements, discussing them, and offering insight or alternatives on how the limitations of the study can be mitigated. This enhances the process of the research and helps readers gain a comprehensive understanding of a study’s conditions.
- Identify the research constraints : Identify those limitations having the greatest impact on the quality of the research findings and your ability to effectively answer your research questions and/or hypotheses. These include sample size, selection bias, measurement error, or other issues affecting the validity and reliability of your research.
- Describe their impact on your research : Reflect on the nature of the identified limitations and justify the choices made during the research to identify the impact of the study’s limitations on the research outcomes. Explanations can be offered if needed, but without being defensive or exaggerating them. Provide context for the limitations of your research to understand them in a broader context. Any specific limitations due to real-world considerations need to be pointed out critically rather than justifying them as done by some other author group or groups.
- Mention the opportunity for future investigations : Suggest ways to overcome the limitations of the present study through future research. This can help readers understand how the research fits into the broader context and offer a roadmap for future studies.
Frequently Asked Questions
- Should I mention all the limitations of my study in the research report?
Restrict limitations to what is pertinent to the research question under investigation. The specific limitations you include will depend on the nature of the study, the research question investigated, and the data collected.
- Can the limitations of a study affect its credibility?
Stating the limitations of the research is considered favorable by editors and peer reviewers. Connecting your study’s limitations with future possible research can help increase the focus of unanswered questions in this area. In addition, admitting limitations openly and validating that they do not affect the main findings of the study increases the credibility of your study. However, if you determine that your study is seriously flawed, explain ways to successfully overcome such flaws in a future study. For example, if your study fails to acquire critical data, consider reframing the research question as an exploratory study to lay the groundwork for more complete research in the future.
- How can I mitigate the limitations of my study?
Strategies to minimize limitations of the research should focus on convincing reviewers and readers that the limitations do not affect the conclusions of the study by showing that the methods are appropriate and that the logic is sound. Here are some steps to follow to achieve this:
- Use data that are valid.
- Use methods that are appropriate and sound logic to draw inferences.
- Use adequate statistical methods for drawing inferences from the data that studies with similar limitations have been published before.
Admit limitations openly and, at the same time, show how they do not affect the main conclusions of the study.
- Can the limitations of a study impact its publication chances?
Limitations in your research can arise owing to restrictions in methodology or research design. Although this could impact your chances of publishing your research paper, it is critical to explain your study’s limitations to your intended audience. For example, it can explain how your study constraints may impact the results and views generated from your investigation. It also shows that you have researched the flaws of your study and have a thorough understanding of the subject.
- How can limitations in research be used for future studies?
The limitations of a study give you an opportunity to offer suggestions for further research. Your study’s limitations, including problems experienced during the study and the additional study perspectives developed, are a great opportunity to take on a new challenge and help advance knowledge in a particular field.
References:
- Brutus, S., Aguinis, H., & Wassmer, U. (2013). Self-reported limitations and future directions in scholarly reports: Analysis and recommendations. Journal of Management , 39 (1), 48-75.
- Ioannidis, J. P. (2007). Limitations are not properly acknowledged in the scientific literature. Journal of Clinical Epidemiology , 60 (4), 324-329.
- Price, J. H., & Murnan, J. (2004). Research limitations and the necessity of reporting them. American Journal of Health Education , 35 (2), 66.
- Boddy, C. R. (2016). Sample size for qualitative research. Qualitative Market Research: An International Journal , 19 (4), 426-432.
R Discovery is a literature search and research reading platform that accelerates your research discovery journey by keeping you updated on the latest, most relevant scholarly content. With 250M+ research articles sourced from trusted aggregators like CrossRef, Unpaywall, PubMed, PubMed Central, Open Alex and top publishing houses like Springer Nature, JAMA, IOP, Taylor & Francis, NEJM, BMJ, Karger, SAGE, Emerald Publishing and more, R Discovery puts a world of research at your fingertips.
Try R Discovery Prime FREE for 1 week or upgrade at just US$72 a year to access premium features that let you listen to research on the go, read in your language, collaborate with peers, auto sync with reference managers, and much more. Choose a simpler, smarter way to find and read research – Download the app and start your free 7-day trial today !
Related Posts

Article Processing Charges: Impact on Open Access Publishing

How Publishers Can Enhance Reader Engagement with R Discovery’s Article Recommendation System
Limitations of Research
- January 2017
- In book: The SAGE Encyclopedia of Communication Research Methods (pp.863-864)
- Publisher: SAGE Publications, Inc

- University of Oklahoma
Discover the world's research
- 25+ million members
- 160+ million publication pages
- 2.3+ billion citations
- INFORM SYST

- Recruit researchers
- Join for free
- Login Email Tip: Most researchers use their institutional email address as their ResearchGate login Password Forgot password? Keep me logged in Log in or Continue with Google Welcome back! Please log in. Email · Hint Tip: Most researchers use their institutional email address as their ResearchGate login Password Forgot password? Keep me logged in Log in or Continue with Google No account? Sign up
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
Preview improvements coming to the PMC website in October 2024. Learn More or Try it out now .
- Advanced Search
- Journal List
- v.28(1); Jan-Mar 2024
- PMC10882193

Limitations in Medical Research: Recognition, Influence, and Warning
Douglas e. ott.
Mercer University, Macon, Georgia, USA.
Background:
As the number of limitations increases in a medical research article, their consequences multiply and the validity of findings decreases. How often do limitations occur in a medical article? What are the implications of limitation interaction? How often are the conclusions hedged in their explanation?
To identify the number, type, and frequency of limitations and words used to describe conclusion(s) in medical research articles.
Search, analysis, and evaluation of open access research articles from 2021 and 2022 from the Journal of the Society of Laparoscopic and Robotic Surgery and 2022 Surgical Endoscopy for type(s) of limitation(s) admitted to by author(s) and the number of times they occurred. Limitations not admitted to were found, obvious, and not claimed. An automated text analysis was performed for hedging words in conclusion statements. A limitation index score is proposed to gauge the validity of statements and conclusions as the number of limitations increases.
A total of 298 articles were reviewed and analyzed, finding 1,764 limitations. Four articles had no limitations. The average was between 3.7% and 6.9% per article. Hedging, weasel words and words of estimative probability description was found in 95.6% of the conclusions.
Conclusions:
Limitations and their number matter. The greater the number of limitations and ramifications of their effects, the more outcomes and conclusions are affected. Wording ambiguity using hedging or weasel words shows that limitations affect the uncertainty of claims. The limitation index scoring method shows the diminished validity of finding(s) and conclusion(s).
INTRODUCTION
As the number of limitations in a medical research article increases, does their influence have a more significant effect than each one considered separately, making the findings and conclusions less reliable and valid? Limitations are known variables that influence data collection and findings and compromise outcomes, conclusions, and inferences. A large body of work recognizes the effect(s) and consequence(s) of limitations. 1 – 77 Other than the ones known to the author(s), unknown and unrecognized limitations influence research credibility. This study and analysis aim to determine how frequently and what limitations are found in peer-reviewed open-access medical articles for laparoscopic/endoscopic surgeons.
This research is about limitations, how often they occur and explained and/or justified. Failure to disclose limitations in medical writing limits proper decision-making and understanding of the material presented. All articles have limitations and constraints. Not acknowledging limitations is a lack of candor, ignorance, or a deliberate omission. To reduce the suspicion of invalid conclusions limitations and their effects must be acknowledged and explained. This allows for a clearer more focused assessment of the article’s subject matter without explaining its findings and conclusions using hedging and words of estimative probability. 78 , 79
An evaluation of open access research/meta-analysis/case series/methodologies/review articles published in the Journal of the Society of Laparoendoscopic and Robotic Surgery ( JSLS ) for 2021 and 2022 (129) and commentary/guidelines/new technology/practice guidelines/review/SAGES Masters Program articles in Surgical Endoscopy ( Surg Endosc ) for 2022 (169) totaling 298 were read and evaluated by automated text analysis for limitations admitted to by the paper’s authors using such words as “limitations,” “limits,” “shortcomings,” “inadequacies,” “flaws,” “weaknesses,” “constraints,” “deficiencies,” “problems,” and “drawbacks” in the search. Limitations not mentioned were found by reading the paper and assigning type and frequency. The number of hedging and weasel words used to describe the conclusion or validate findings was determined by reading the article and adding them up.
For JSLS , there were 129 articles having 63 different types of limitations. Authors claimed 476, and an additional 32 were found within the article, totaling 508 limitations (93.7% admitted to and 6.3% discovered that were not mentioned). This was a 3.9 limitation average per article. No article said it was free of limitations. The ten most frequent limitations and their rate of occurrence are in Table 1 . The total number of limitations, frequency, and visual depictions are seen in Figures 1A and and 1B 1B .

( A ) Visual depiction of the ranked frequency of limitations for JSLS articles reviewed.
The Ten Most Frequent Limitations Found in JSLS and Surg Endosc Articles
| top 10 limitations | Total number of limitations | Number of articles | Percent of total number of limitations | top 10 limitations | Total number of limitations | Number of articles | Percent of total number of limitations |
|---|---|---|---|---|---|---|---|
| Results not generalizable | 33 | 33/508 | 6.5% | Results not generalizable | 86 | 86/1256 | 6.8% |
| Retrospective study | 32 | 32/508 | 6.3% | Selection bias | 83 | 83/1256 | 6.6% |
| Small sample size | 32 | 32/508 | 6.3% | Confounding variables and comorbidities | 72 | 72/1256 | 5.7% |
| Confounding variables and comorbidities | 23 | 23/508 | 4.5% | Retrospective study | 69 | 69/1256 | 5.5% |
| Selection bias | 21 | 21/508 | 4.1% | Small sample size | 63 | 63/1256 | 5.0% |
| Incomplete data | 20 | 20/508 | 3.9% | Incomplete data | 58 | 58/1256 | 4.6% |
| Limited patient selection criteria | 16 | 16/508 | 3.1% | Lack of standardized treatment | 55 | 55/1256 | 4.4% |
| Limited data availability | 16 | 16/508 | 5.1% | Measurement problems | 53 | 53/1256 | 4.2% |
| No long-term follow-up | 15 | 15/508 | 3.0% | Limited analysis | 47 | 47/1256 | 3.7% |
| Reporting errors | 14 | 14/508 | 2.8% | Problems with study design | 39 | 39/1259 | 3.1% |
| 222/508 | 625/1256 |
There were 169 articles for Surg Endosc , with 78 different named limitations the authors claimed for a total of 1,162. An additional 94 limitations were found in the articles, totaling 1,256, or 7.4 per article. The authors explicitly stated 92.5% of the limitations, and an additional 7.5% of additional limitations were found within the article. Five claimed zero limitations (5/169 = 3%). The ten most frequent limitations and their rate of occurrence are in Table 1 . The total number of limitations and frequency is shown in Figures 1A and and 1B 1B .
Conclusions were described in hedged, weasel words or words of estimative probability 95.6% of the time (285/298).
A research hypothesis aims to test the idea about expected relationships between variables or to explain an occurrence. The assessment of a hypothesis with limitations embedded in the method reaches a conclusion that is inherently flawed. What is compromised by the limitation(s)? The result is an inferential study in the presence of uncertainty. As the number of limitations increases, the validity of information decreases due to the proliferation of uncertain information. Information gathered and conclusions made in the presence of limitations can be functionally unsound. Hypothesis testing of spurious conditions with limitations and then claiming a conclusion is not a reliable method for generating factual evidence. The authors’ reliance on limitation gathered “evidence” data and asserting that this is valid is spurious reasoning. The bridge between theory and evidence is not through limitations that unquestionably accept findings. A range of conclusion possibilities exists being some percent closer to either more correct or incorrect. Relying on leveraging the pursuit of “fact” in the presence of limitations as the safeguard is akin to the fox watching the hen house. Acknowledgment of the uncertainty limitations create in research and discounting the finding’s reliability would give more credibility to the effort. Shortcomings and widespread misuses of research limitation justifications make findings suspect and falsely justified in many instances.
The JSLS instructions to authors say that in the discussion section of the paper the author(s) must “Comment on any methodological weaknesses of the study” ( http://jsls.sls.org/guidelines-for-authors/ ). In their instructions for authors, Surg Endosc says that in the discussion of the paper, “A paragraph discussing study limitations is required” ( https://www.springer.com/journal/464/submission-guidelines ). A comment for a written article about a limitation should express an opinion or reaction. A paragraph discussing limitations, especially, if there is more than one, requires just that: a paragraph and discussion. These requirements were not met or enforced by JSLS 86% (111/129) of the time and 92.3% (156/169) for Surg Endosc . This is an error in peer reviewing, not adhering to established research publication best practices, and the journals needing to adhere to their guidelines. The International Committee of Medical Journal Editors, uniform requirements for manuscripts recommends that authors “State the limitations of your study, and explore the implications of your findings for future research and for clinical practice or policy. Discuss the influence or association of variables, such as sex and/or gender, on your findings, where appropriate, and the limitations of the data.” It also says, “describe new or substantially modified methods, give reasons for using them, and evaluate their limitations” and “Include in the Discussion section the implications of the findings and their limitations, including implications for future research” and “give references to established methods, including statistical methods (see below); provide references and brief descriptions for methods that have been published but are not well known; describe new or substantially modified methods, give reasons for using them, and evaluate their limitations.” 65 “Reporting guidelines (e.g., CONSORT, 1 ARRIVE 2 ) have been proposed to promote the transparency and accuracy of reporting for biomedical studies, and they often include discussion of limitations as a checklist item. Although such guidelines have been endorsed by high-profile biomedical journals, and compliance with them is associated with improved reporting quality, 3 adherence remains suboptimal.” 4 , 5
Limitations start in the methodologic design phase of research. They require troubleshooting evaluations from the start to consider what limitations exist, what is known and unknown, where, and how to overcome them, and how they will affect the reasonableness and assessment of possible conclusions. A named limitation represents a category with numerous components. Each factor has a unique effect on findings and collectively influences conclusion assessment. Even a single limitation can compromise the study’s implementation and adversely influence research parameters, resulting in diminished value of the findings, outcomes, and conclusions. This becomes more problematic as the number of limitations and their components increase. Any limitation influences a research paper. It is unknown how much and to what extent any limitation affects other limitations, but it does create a cascading domino effect of ever-increasing interactions that compromise findings and conclusions. Considering “research” as a system, it has sensitivity and initial conditions (methodology, data collection, analysis, etc.). The slightest alteration of a study due to limitations can profoundly impact all aspects of the study. The presence and influence of limitations introduce a range of unpredictable influences on findings, results, and conclusions.
Researchers and readers need to pay attention to and discount the effects limitations have on the validity of findings. Richard Feynman said in “Cargo cult science” “the first principle is that you must not fool yourself and you are the easiest person to fool.” 73 We strongly believe our own nonsense or wrong-headed reasoning. Buddhist philosophers say we are attached to our ignorance. Researchers are not critical enough about how they fool themselves regarding their findings with known limitations and then pass them on to readers. The competence of findings with known limitations results in suspect conclusions.
Authors should not ask for dismissal, disregard, or indulgence of their limitations. They should be thoughtful and reflective about the implications and uncertainty the limitations create 67 ; their uncertainties, blind spots, and impact on the research’s relevance. A meaningful presentation of study limitations should describe the limitation, explain its effect, provide possible alternative approaches, and describe steps taken to mitigate the limitation. This was largely absent from the articles reviewed.
Authors use synonyms and phrases describing limitations that hide, deflect, downplay, and divert attention from them, i.e., some drawbacks of the study are …, weaknesses of the study are…, shortcomings are…, and disadvantages of the study are…. They then say their finding(s) lack(s) generalizability, meaning the findings only apply to the study participants or that care, sometimes extreme, must be taken in interpreting the results. Which limitation components are they referring to? Are the authors aware of the extent of their limitations, or are they using convenient phrases to highlight the existence of limitations without detailing their defects?
Limitations negatively weigh on both data and conclusions yet no literature exists to provide a quantifiable measure of this effect. The only acknowledgment is that limitations affect research data and conclusions. The adverse effects of limitations are both specific and contextual to each research article and is part of the parameters that affect research. All the limitations are expressed in words, excuses, and a litany of mea culpas asking for forgiveness and without explaining the extent or magnitude of their impact. It is left to the writer and reader to figure out. It is not known what value writers put on their limitations in the 298 articles reviewed from JSLS and Surg Endosc . Listing limitations without comment and effect on the findings and conclusions is a compromising red flag. Therefore, a limitation scoring method was developed and is proposed to assess the level of suspicion generated by the number of limitations.
It is doubtful that a medical research article is so well designed and executed that there are no limitations. This is doubtful since there are unknown unknowns. This study showed that authors need to acknowledge all the limitations when they are known. They acknowledge the ones they know but do not consider other possibilities. There are the known known limitations; the ones the author(s) are aware of and can be measured, some explained, most not. The known unknowns: limitations authors are aware of but cannot explain or quantify. The unknown unknown limitations: the ones authors are not aware of and have unknown influence(s), i.e., the things they do not know they do not know. These are blind spots (not knowing what they do not know or black swan events). And the unknown knowns; the limitations authors may be aware of but have not disclosed, thoroughly reported, understood, or addressed. They are unexpected and not considered. See Table 2 . 74
Limitations of Known and Unknowns as They Apply to Limitations
| Things we are aware of and understand. | Things we are aware of but don’t understand. | |
| Things we understand but are not aware of. | Things we are neither aware of nor understand. | |
It is possible that authors did not identify, want to identify, or acknowledge potential limitations or were unaware of what limitations existed. Cumulative complexity is the result of the presence of multiple limitations because of the accumulation and interaction of limitations and their components. Just mentioning a limitation category and not the specific parts that are the limitation(s) is not enough. Authors telling readers of their known research limitations is a caution to discount the findings and conclusions. At what point does the caution for each limitation, its ramifications, and consequences become a warning? When does the piling up of mistakes, bad and missing data, biases, small sample size, lack of generalizability, confounding factors, etc., reach a point when the findings become s uninterpretable and meaningless? “Caution” indicates a level of potential hazard; a warning is more dire and consequential. Authors use the word “caution” not “warning” to describe their conclusions. There is a point when the number of limitations and their cumulative effects surpasses the point where a caution statement is no longer applicable, and a warning statement is required. This is the reason for establishing a limitations risk score.
Limitations put medical research articles at risk. The accumulation of limitations (variables having additional limitation components) are gaps and flaws diluting the probability of validity. There is currently no assessment method for evaluating the effect(s) of limitations on research outcomes other than awareness that there is an effect. Authors make statements warning that their results may not be reliable or generalizable, and need more research and larger numbers. Just because the weight effect of any given limitation is not known, explained, or how it discounts findings does not negate a causation effect on data, its analysis, and conclusions. Limitation variables and the ramifications of their effects have consequences. The relationship is not zero effect and accumulates with each added limitation.
As a result of this research, a limitation index score (LIS) system and assessment tool were developed. This limitation risk assessment tool gives a scores assessment of the relative validity of conclusions in a medical article having limitations. The adoption of the LIS scoring assessment tool for authors, researchers, editors, reviewers, and readers is a step toward understanding the effects of limitations and their causal relationships to findings and conclusions. The objective is cleaner, tighter methodologies, and better data assessment, to achieve more reliable findings. Adjustments to research conclusions in the presence of limitations are necessary. The degree of modification depends on context. The cumulative effect of this burden must be acknowledged by a tangible reduction and questioning of the legitimacy of statements made under these circumstances. The description calculating the LIS score is detailed in Appendix 1 .
A limitation word or phrase is not one limitation; it is a group of limitations under the heading of that word or phrase having many additional possible components just as an individual named influence. For instance, when an admission of selection bias is noted, the authors do not explain if it was an exclusion criterion, self-selection, nonresponsiveness, lost to follow-up, recruitment error, how it affects external validity, lack of randomization, etc., or any of the least 263 types of known biases causing systematic distortions of the truth whether unintentional or wanton. 40 , 76 Which forms of selection bias are they identifying? 63 Limitations have branches that introduce additional limitations influencing the study’s ability to reach a useful conclusion. Authors rarely tell you the effect consequences and extent limitations have on their study, findings, and conclusions.
This is a sample of limitations and a few of their component variables under the rubric of a single word or phrase. See Table 3 .
A Limitation Word or Phrase is a Limitation Having Additional Components That Are Additional Limitations. When an Author Uses the Limitation Composite Word or Phrase, They Leave out Which One of Its Components is Contributory to the Research Limitations. Each Limitation Interacts with Other Limitations, Creating a Cluster of Cross Complexities of Data, Findings, and Conclusions That Are Tainted and Negatively Affect Findings and Conclusions
| Small Sample Size | Retrospective Study | Selection Bias |
|---|---|---|
| Low statistical power | Missing information | Affects internal validity |
| Estimates not reliable | Recall bias | Nonrandom selection |
| Prone to biased samples | Observer bias | Leads to confounding |
| Not generalizable | Misclassification bias | Not generalizable |
| Prone to false negative error | Observer bias | Inaccurate relation to variables |
| Prone to false positive error | Evidence less robust than prospective study | Observer bias |
| Sampling error | Missing data | Sampling bias |
| Confounding factors | Volunteer bias | |
| Selection bias | Survivorship bias |
Limitations rarely occur alone. If you see one there are many you do not see or appreciate. Limitation s components interact with their own and other limitations, leading to complex connections interacting and discounting the reliability of findings. By how much is context dependent: but it is not zero. Limitations are variables influencing outcomes. As the number of limitations increases, the reliability of the conclusions decreases. How many variables (limitations) does it take to nullify the claims of the findings? The weight and influence of each limitation, its aggregate components, and interconnectedness have an unknown magnitude and effect. The result is a disorderly concoction of hearsay explanations. Table 4 is an example of just two single explanation limitations and some of their components illustrating the complex compounding of their effects on each other.
An Example of Interactions between Only Two Limitations and Some of Their Components Causes 16 Interactions
| Retrospective Study | Small Sample Size |
|---|---|
The novelty of this paper on limitations in medical science is not the identification of research article limitations or their number or frequency; it is the recognition of the multiplier effect(s) limitations and the influence they have on diminishing any conclusion(s) the paper makes. It is possible that limitations contribute to the inability of studies to replicate and why so many are one-time occurrences. Therefore, the generalizability statement that should be given to all readers is BEWARE THERE IS A REDUCTION EFFECT ON THE CONCLUSIONS IN THIS ARTICLE BECAUSE OF ITS LIMITATIONS.
Journals accept studies done with too many limitations, creating forking path situations resulting in an enormous number of possible associations of individual data points as multiple comparisons. 79 The result is confusion, a muddled mess caused by interactions of limitations undermining the ability to make valid inferences. Authors know and acknowledge but rarely explain them or their influence. They also use incomplete and biased databases, biased methods, small sample sizes, and not eliminating confounders, etc., but persist in doing research with these circumstances. Why is that? Is it because when limitations are acknowledged, authors feel justified in their conclusions? It wasn’t my poor research design; it was the limitation(s). How do peer reviewers score and analyze these papers without a method to discount the findings and conclusions in the presence of limitations? What are the calculus editors use to justify papers with multiple limitations, reaching compromised or spurious conclusions? How much caution or warning should a journal say must be taken in interpreting article results? How much? Which results? When? Under what circumstance(s)?
Since a critical component of research is its limitations, the quality and rigor of research are largely defined by, 75 these constraints making it imperative that limitations be exposed and explained. All studies have limitations admitted to or not, and these limitations influence outcomes and conclusions. Unfortunately, they are given insufficient attention, accompanied by feeble excuses, but they all matter. The degrees of freedom of each limitation influence every other limitation, magnifying their ramifications and confusion. Limitations of a scientific article must put the findings in context so the reader can judge the validity and strength of the conclusions. While authors acknowledge the limitations of their study, they influence its legitimacy.
Not only are limitations not properly acknowledged in the scientific literature, 8 but their implications, magnitude, and how they affect a conclusion are not explained or appreciated. Authors work at claiming their work and methods “overcome,” “avoid,” or “circumvent” limitations. Limitations are explained away as “Failure to prove a difference does not prove lack of a difference.” 60 Sample size, bias, confounders, bad data, etc. are not what they seem and do not sully the results. The implication is “trust me.” But that’s not science. Limitations create cognitive distortions and framing (misperception of reality) for the authors and readers. Data in studies with limitations is data having limitations. It was real but tainted.
Limitations are not a trivial aspect of research. It is a tangible something, positive or negative, put into a data set to be analyzed and used to reach a conclusion. How did these extra somethings, known unknowns, not knowns, and unknown knowns, affect the validity of the data set and conclusions? Research presented with the vagaries of explicit limitations is intensified by additional limitations and their component effects on top of the first limitation s , quickly diluting any conclusion making its dependability questionable.
This study’s analysis of limitations in medical articles averaged 3.9% per article for JSLS and 7.4% for Surg Endosc . Authors admit to some and are aware of limitations, but not all of them and discount or leave out others. Limitations were often presented with misleading and hedging language. Authors do not give weight or suggest the percent discount limitations have on the reliance of conclusion(s). Since limitations influence findings, reliability, generalizability, and validity without knowing the magnitude of each and their context, the best that can be said about the conclusions is that they are specific to the study described, context-driven, and suspect.
Limitations mean something is missing, added, incorrect, unseen, unaware of, fabricated, or unknown; circumstances that confuse, confound, and compromise findings and information to the extent that a notice is necessary. All medical articles should have this statement, “Any conclusion drawn from this medical study should be interpreted considering its limitations. Readers should exercise caution, use critical judgement, and consult other sources before accepting these findings. Findings may not be generalizable regardless of sample size, composition, representative data points, and subject groups. Methodologic, analytic, and data collection may have introduced biases or limitations that can affect the accuracy of the results. Controlling for confounding variables, known and unknown, may have influenced the data and/or observations. The accuracy and completeness of the data used to draw a conclusion may not be reliable. The study was specific to time, place, persons, and prevailing circumstances. The weight of each of these factors is unknown to us. Their effect may be limited or compounded and diminish the validity of the proposed conclusions.”
This study and findings are limited and constrained by the limitations of the articles reviewed. They have known and unknown limitations not accounted for, missing data, small sample size, incongruous populations, internal and external validity concerns, confounders, and more. See Tables 2 and and 3 . 3 . Some of these are correctible by the author’s awareness of the consequences of limitations, making plans to address them in the methodology phase of hypothesis assessment and performance of the research to diminish their effects.
Limitations in research articles are expected, but they can be reduced in their effect so that conclusions are closer to being valid. Limitations introduce elements of ignorance and suspicion. They need to be explained so their influence on the believability of the study and its conclusions is closer to meeting construct, content, face, and criterion validity. As the number of limitations increases, common sense, skepticism, study component acceptability, and understanding the ramifications of each limitation are necessary to accept, discount, or reject the author’s findings. As the number of hedging and weasel words used to explain conclusion(s) increases, believability decreases, and raises suspicion regarding claims. Establishing a systematic limitation scoring index limitations for authors, editors, reviewers, and readers and recognizing their cumulative effects will result in a clearer understanding of research content and legitimacy.
How to calculate the Limitation Index Score (LIS). See Tables 5 – 5 . Each limitation admitted to by authors in the article equals (=) one (1) point. Limitations may be generally stated by the author as a broad category, but can have multiple components, such as a retrospective study with these limitation components: 1. data or recall not accurate, 2. data missing, 3. selection bias not controlled, 4. confounders not controlled, 5. no randomization, 6. no blinding, 7. difficult to establish cause and effect, and 8. cannot draw a conclusion of causation. For each component, no matter how many are not explained and corrected, add an additional one (1) point to the score. See Table 2 .
The Limitation Scoring Index is a Numeric Limitation Risk Assessment Score to Rank Risk Categories and Discounting Probability of Validity and Conclusions. The More Limitations in a Study, the Greater the Risk of Unreliable Findings and Conclusions
| Number of limitations | Word description of discounting | Proposed percent discounting of conclusions | Outcome probability | Increasing level of less reliable conclusions |
|---|---|---|---|---|
| 0 | Unknown unknowns | 1–10% | May have valid conclusion(s) | Warning |
| 1–2 | Some | 15–25% | ↓ | ↓ |
| 3–4 | Probable | 35–45% | ↓ | Caution |
| 5–6 | Likely | 70–80% | ↓ | ↓ |
| 7–8 | Highly likely | 85–95% | ↓ | ↓ |
| >8 | Certain | 97–100% | Very questionable conclusion(s) | Danger |
Limitations May Be Generally Stated by the Author but Have Multiple Components, Such as a Retrospective Study Having Disadvantage Components of 1. Data or Recall Not Accurate, 2. Data Missing, 3. Selection Bias Not Controlled, 4. Confounders Not Controlled, 5. No Randomization, 6. No Blinding, 7 Difficult to Establish Cause and Effect, 8. Results Are Hypothesis Generating, and 9. Cannot Draw a Conclusion of Causation. For Each Component, Not Explained and Corrected, Add an Additional One (1) Point Is Added to the Score. Extra Blanks Are for Additional Limitations
| One point for each limitation | |
|---|---|
| One additional point for each component of each limitation | |
| Retrospective study | |
| Small sample size | |
| Not generalizable | |
| Selection bias | |
| Not controlling for confounders | |
| Not controlling for comorbidities | |
| Incomplete or missing data | |
| No long-term follow-up | |
| Reporting errors | |
| Measurement problems | |
| Study design problems | |
| Lack of standardized treatment | |
| Subtotal for Table 2 |
An Automatic 2 Points is Added for Meta-Analysis Studies Since They Have All the Retrospective Detrimental Components. 44 Data from Insurance, State, National, Medicare, and Medicaid, Because of Incorrect Coding, Over Reporting, and Under-Reporting, Etc. Each Component of the Limitation Adds One Additional Point. For Surveys and Questionnaires Add One Additional Point for Each Bias. Extra Blanks Are for Additional Limitations
| Two points for these limitations | |
|---|---|
| One additional point for each limitation and one additional point for each limitation component. | |
| Meta-analysis | |
| Data from Medicare, Medicaid, insurance companies, disease, state, and national databases | |
| Surveys and questionnaires | |
| Each limitation not admitted to | |
| Subtotal for Table 3 |
Automatic Five (5) Points for Manufacturer and User Facility Device Experience (MAUDE) Database Articles. The FDA Access Data Site Says Submissions Can Be “Incomplete, Inaccurate, Untimely, Unverified, or Biased” and “the Incidence or Prevalence of an Event Cannot Be Determined from This Reporting System Alone Due to Under-Reporting of Events, Inaccuracies in Reports, Lack of Verification That the Device Caused the Reported Event, and Lack of Information” and “DR Data Alone Cannot Be Used to Establish Rates of Events, Evaluate a Change in Event Rates over Time or Compare Event Rates between Devices. The Number of Reports Cannot Be Interpreted or Used in Isolation to Reach Conclusions” 80
| Five points for MAUDE based articles | |
|---|---|
| One additional point for each additional limitation and one point for each of its components. | |
| Subtotal for Table 4 |
Total Limitation Index Score
| Limitations | Calculation |
|---|---|
| Subtotal for Table 2 | |
| Subtotal for Table 3 | |
| Subtotal for Table 4 | |
| Total Limitation Index Score |
Each limitation not admitted to = two (2) points. A meta-analysis study gets an automatic 2 points since they are retrospective and have detrimental components that should be added to the 2 points. Data from insurance, state, national, Medicare, and Medicaid, because of incorrect coding, over-reporting, and underreporting, etc., score 2 points, and each component adds one additional point. Surveys and questionnaires get 2 points, and add one additional point for each bias. See Table 3 .
Manufacturer and User Facility Device Experience (MAUDE) database articles receive an automatic five (5) points. The FDA access data site says, submissions can be “incomplete, inaccurate, untimely, unverified, or biased” and “the incidence or prevalence of an event cannot be determined from this reporting system alone due to underreporting of events, inaccuracies in reports, lack of verification that the device caused the reported event, and lack of information” and “MDR data alone cannot be used to establish rates of events, evaluate a change in event rates over time or compare event rates between devices. The number of reports cannot be interpreted or used in isolation to reach conclusions.” 80 See Table 4 . Add one additional point for each additional limitation noted in the article.
Add one additional point for each additional limitation and one point for each of its components. Extra blanks are for additional
limitations and their component scores.
Funding sources: none.
Disclosure: none.
Conflict of interests: none.
Acknowledgments: Author would like to thank Lynda Davis for her help with data collection.
References:
All references have been archived at https://archive.org/web/
When you choose to publish with PLOS, your research makes an impact. Make your work accessible to all, without restrictions, and accelerate scientific discovery with options like preprints and published peer review that make your work more Open.
- PLOS Biology
- PLOS Climate
- PLOS Complex Systems
- PLOS Computational Biology
- PLOS Digital Health
- PLOS Genetics
- PLOS Global Public Health
- PLOS Medicine
- PLOS Mental Health
- PLOS Neglected Tropical Diseases
- PLOS Pathogens
- PLOS Sustainability and Transformation
- PLOS Collections
- How to Write Discussions and Conclusions

The discussion section contains the results and outcomes of a study. An effective discussion informs readers what can be learned from your experiment and provides context for the results.
What makes an effective discussion?
When you’re ready to write your discussion, you’ve already introduced the purpose of your study and provided an in-depth description of the methodology. The discussion informs readers about the larger implications of your study based on the results. Highlighting these implications while not overstating the findings can be challenging, especially when you’re submitting to a journal that selects articles based on novelty or potential impact. Regardless of what journal you are submitting to, the discussion section always serves the same purpose: concluding what your study results actually mean.
A successful discussion section puts your findings in context. It should include:
- the results of your research,
- a discussion of related research, and
- a comparison between your results and initial hypothesis.
Tip: Not all journals share the same naming conventions.
You can apply the advice in this article to the conclusion, results or discussion sections of your manuscript.
Our Early Career Researcher community tells us that the conclusion is often considered the most difficult aspect of a manuscript to write. To help, this guide provides questions to ask yourself, a basic structure to model your discussion off of and examples from published manuscripts.
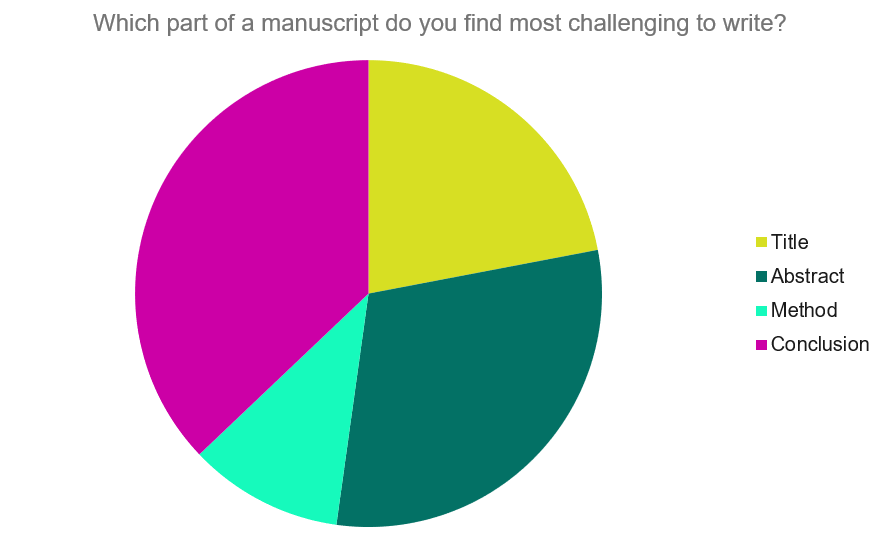
Questions to ask yourself:
- Was my hypothesis correct?
- If my hypothesis is partially correct or entirely different, what can be learned from the results?
- How do the conclusions reshape or add onto the existing knowledge in the field? What does previous research say about the topic?
- Why are the results important or relevant to your audience? Do they add further evidence to a scientific consensus or disprove prior studies?
- How can future research build on these observations? What are the key experiments that must be done?
- What is the “take-home” message you want your reader to leave with?
How to structure a discussion
Trying to fit a complete discussion into a single paragraph can add unnecessary stress to the writing process. If possible, you’ll want to give yourself two or three paragraphs to give the reader a comprehensive understanding of your study as a whole. Here’s one way to structure an effective discussion:
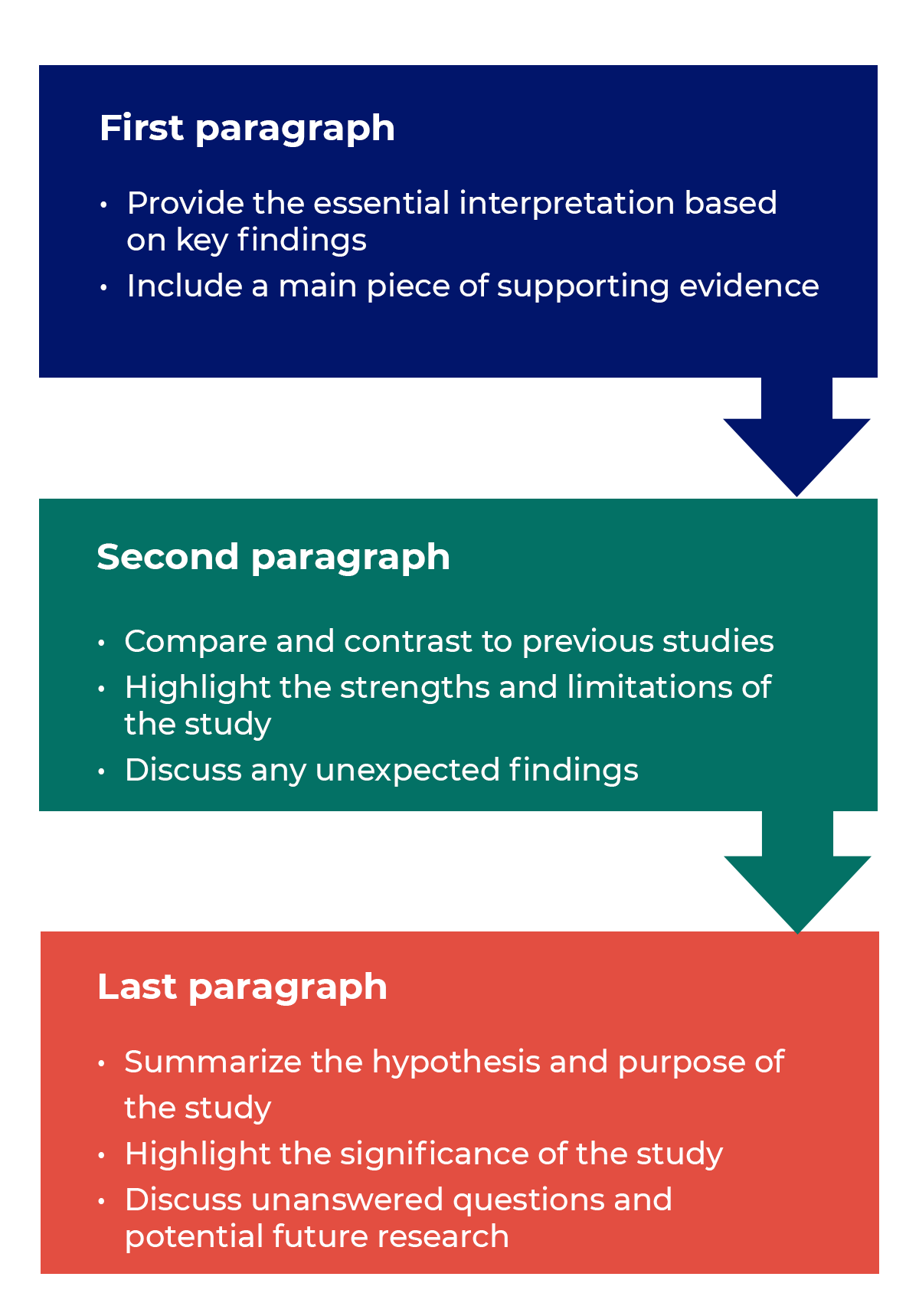
Writing Tips
While the above sections can help you brainstorm and structure your discussion, there are many common mistakes that writers revert to when having difficulties with their paper. Writing a discussion can be a delicate balance between summarizing your results, providing proper context for your research and avoiding introducing new information. Remember that your paper should be both confident and honest about the results!

- Read the journal’s guidelines on the discussion and conclusion sections. If possible, learn about the guidelines before writing the discussion to ensure you’re writing to meet their expectations.
- Begin with a clear statement of the principal findings. This will reinforce the main take-away for the reader and set up the rest of the discussion.
- Explain why the outcomes of your study are important to the reader. Discuss the implications of your findings realistically based on previous literature, highlighting both the strengths and limitations of the research.
- State whether the results prove or disprove your hypothesis. If your hypothesis was disproved, what might be the reasons?
- Introduce new or expanded ways to think about the research question. Indicate what next steps can be taken to further pursue any unresolved questions.
- If dealing with a contemporary or ongoing problem, such as climate change, discuss possible consequences if the problem is avoided.
- Be concise. Adding unnecessary detail can distract from the main findings.

Don’t
- Rewrite your abstract. Statements with “we investigated” or “we studied” generally do not belong in the discussion.
- Include new arguments or evidence not previously discussed. Necessary information and evidence should be introduced in the main body of the paper.
- Apologize. Even if your research contains significant limitations, don’t undermine your authority by including statements that doubt your methodology or execution.
- Shy away from speaking on limitations or negative results. Including limitations and negative results will give readers a complete understanding of the presented research. Potential limitations include sources of potential bias, threats to internal or external validity, barriers to implementing an intervention and other issues inherent to the study design.
- Overstate the importance of your findings. Making grand statements about how a study will fully resolve large questions can lead readers to doubt the success of the research.
Snippets of Effective Discussions:
Consumer-based actions to reduce plastic pollution in rivers: A multi-criteria decision analysis approach
Identifying reliable indicators of fitness in polar bears
- How to Write a Great Title
- How to Write an Abstract
- How to Write Your Methods
- How to Report Statistics
- How to Edit Your Work
The contents of the Peer Review Center are also available as a live, interactive training session, complete with slides, talking points, and activities. …
The contents of the Writing Center are also available as a live, interactive training session, complete with slides, talking points, and activities. …
There’s a lot to consider when deciding where to submit your work. Learn how to choose a journal that will help your study reach its audience, while reflecting your values as a researcher…
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
- Review Article
- Open access
- Published: 17 June 2024
Limitations of current techniques in clinical antimicrobial resistance diagnosis: examples and future prospects
- Jack Hassall 1 ,
- Carmen Coxon 1 ,
- Vishal C. Patel 2 , 3 , 4 ,
- Simon D. Goldenberg 5 &
- Chrysi Sergaki ORCID: orcid.org/0000-0003-0477-8680 1
npj Antimicrobials and Resistance volume 2 , Article number: 16 ( 2024 ) Cite this article
226 Accesses
1 Altmetric
Metrics details
- Antimicrobial resistance
- Health services
- Infectious-disease diagnostics
- Next-generation sequencing
Antimicrobial resistance is a global threat to public health. Without proactive intervention, common infections may become untreatable, restricting the types of clinical intervention that can be undertaken and reversing improvements in mortality rates. Effective antimicrobial stewardship represents one approach to restrict the spread of antimicrobial resistance but relies on rapid and accurate diagnostics that minimise the unnecessary use of antibiotics. This is increasingly a key unmet clinical need. In this paper, we describe existing techniques for the detection of antimicrobial resistance, while examining their drawbacks and limitations. We also discuss emerging diagnostic technologies in the field, and the need for standardisation to allow for swifter and more widespread clinical adoption.
Similar content being viewed by others
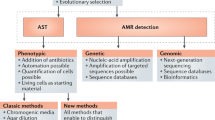
Innovative and rapid antimicrobial susceptibility testing systems
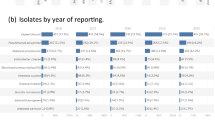
A nested cohort 5-year Canadian surveillance of Gram-negative antimicrobial resistance for optimized antimicrobial therapy

ESKAPE pathogens: antimicrobial resistance, epidemiology, clinical impact and therapeutics
Introduction.
Since the introduction of penicillin during World War II, antibiotics have become the backbone of modern medicine 1 , 2 . The success of antibiotics resulted in a golden age in medicine, but this is now coming to an end as we risk entering a post-antibiotic era. A lack of effective antibiotics reduces our capacity to respond to outbreaks of infectious disease. Without coordinated, proactive interventions to detect and manage antimicrobial resistance (AMR), there will be a significant regression in medical care and a steep increase in mortality rates 3 . Many modern medical techniques are dependent on the availability of effective antimicrobials, without them many common procedures and interventions (cancer chemotherapy, organ transplantation, prosthetic joint replacement etc) may not be able to be undertaken without excess risk 3 .
Multi-drug resistant organisms (MDROs) are the outcome of years of antibiotic dependency in medical practice and are responsible for an increasing number of infections. MDRO are categorised by three increasing resistance levels 4 :
Multidrug-resistant (MDR) – nonsusceptibility to at least one agent in three or more antimicrobial agent classes.
Extensively drug-resistant (XDR) – nonsusceptibility to at least one agent in all but two (or fewer) antimicrobial agent classes.
Pan drug-resistant (PDR) or sometimes referred to as totally drug-resistant (TDR) whereby the organism shows nonsusceptibility to all agents in all classes.
MDROs are considered a global crisis affecting low, middle and high-income countries 5 , given their potentially untreatable nature 6 . AMR does not respect borders, neither geographical nor ecological, and with the use of antimicrobials in food producing animals we are seeing transmission of resistant pathogens from livestock into humans 7 . In 2015, the Global Action Plan on Antimicrobial Resistance was established by the World Health Assembly to address the threat of AMR 5 , 8 . This was followed by the United Nations General Assembly that passed a resolution unanimously calling for a globally coordinated action resulting in the One Health approach to AMR 5 , 9 , 10 . The One Health approach is a multidisciplinary joint effort to provide solutions for human, animal, and environmental health 11 .
The socioeconomic burden of AMR is difficult to gauge but known to be significant. Mortality estimates range from 0.7–4.95 million deaths worldwide annually, and healthcare costs amounting to tens of billions of US dollars 8 , 10 , 12 , 13 , 14 . It is likely that these numbers are an underestimate due to insufficient national reporting rates, a lack of comprehensive data coverage and no International Classification of Diseases (ICD-10) code specifically for MDRO infections 15 . In 2018, Burnham et al. re-analysed the 2010 data for MDRO-related deaths in the US, identifying 154,113 deaths vs an original estimation of 23,000, nearly seven times the original CDC estimate 15 . What we do know, is that MDROs are on the rise, with the number of reported MDR strains quadrupling over the last two decades, particularly in young children, accounting for 5–10% hospitalised cases 16 , 17 , 18 .
With the world population now around 8 billion and over 55% of all people concentrated in densely populated urban centres 19 ; the risk of a bacterial pandemic is increasingly likely without effective control 20 . The COVID-19 pandemic has been a stark reminder of the ferocity at which an infectious disease can spread and the extensive damage it can cause 21 . Surveillance of infectious agents must improve to allow us to better prepare for and limit future outbreaks, reducing our dependency on antibiotics. Furthermore, tracking how, where, and at what rate antibiotic resistance is evolving in bacteria, can aid in predicting and fighting outbreaks of AMR infections. Currently, patients with suspected infections are most likely to be treated empirically, with some countries estimating 30–50% inappropriate or unnecessary antibiotic usage 22 . This is disappointing considering the significant progress that has been made towards fast, accurate, and affordable diagnostics and the availability of antimicrobial resistance screening.
Methods used in AMR tracking
The European Committee on Antimicrobial Susceptibility Testing (EUCAST) and the Clinical Laboratory Standards Institute (CLSI) recommend investigating bacterial resistance to antibiotics using culture-based techniques 23 , 24 , 25 : the current gold standard for verifying AMR. Culture-based assessment involves observing and reporting the growth (or absence of growth) of bacteria exposed to various concentrations of antibiotics (Table 1 ). Culture-based approaches can be used to establish a minimum inhibitory concentration (MIC) or minimum bactericidal concentration (MBC) for a particular organism-antibiotic combination, giving an indication of the likelihood that a particular agent will be clinically effective. The main advantage of assessing AMR this way is the low cost, as the consumables and equipment are inexpensive compared to PCR 26 , 27 . However, some scientists argue that this labour intensive and slow approach is too costly both in terms of laboratory staff costs and extended in-patient times 26 . Less labour-intensive culture methods do exist, such as Disk and Strip diffusion gradient, but these are still time-costly and laborious to perform when testing multiple samples and antibiotics.
Culture-based assessment relies on the ability to isolate the strain of interest from a complex mixture, and it is also essential that the species is compatible with the culturing technique (e.g. anaerobic bacteria cannot grow in normal atmospheric conditions). To identify strains, a sample is initially grown on solid media and any colonies that form can be identified using a variety of techniques; amplification and sequencing based (16 S and PCR), biochemically (Analytical profile index, API), immunologically (Enzyme-linked immunosorbent assay, ELISA), or through protein fragment analysis using matrix-assisted laser desorption/ionisation time-of-flight mass spectrophotometry (MALDI-TOF MS) 28 . Once the species is/are identified, the antimicrobial susceptibility can be determined. Complex and non-sterile sample types such as faeces make the culture-based assessment difficult, as a plethora of colonies will grow during the initial culturing step. Selective media can be used in this case to target recovery of a suspected microbe of concern 29 .
Lateral flow tests (LFTs) can be used in the context of AMR assessment, the technology uses an immunochromatographic strip impregnated with antibodies to detect key enzymes associated with antimicrobial resistance e.g. beta-lactamase. LFTs proved highly successful during the COVID-19 pandemic 30 , 31 . However, LFTs have limited use in the context of AMR testing, as there is currently a requirement to first undertake pre-culture step 24 . So, while quick, LFTs are still limited by bacterial growth times and the capacity to undertake these steps 32 .
Molecular techniques for pathogen detection and antibiotic resistance mechanisms are an attractive alternative to culture-based methods due to their high selectivity at the RNA/DNA level, sensitivity, and ability to provide earlier identification (or diagnosis) 24 , 33 , 34 . While molecular methods are more expensive than culture-based ones (cost per test), one could argue that the benefit of earlier diagnosis, patient discharge from hospital, and fewer working days lost, presents cost savings in the wider context 26 .
Nucleic acid amplification tests (NAATs) for detection of pathogens can use a variety of amplification methods (PCR, Strand Displacement Amplification (SDA), Transcription-Mediated Amplification), but are mainly limited to PCR for antibiotic resistance gene detection 24 . PCR species identification is highly targeted and requires a level of empirical insight from medical professionals to narrow down the range of causative agents to direct screening. This is also true for AMR – the mechanism of resistance in the pathogen of interest must be known to allow for the design of targeted PCR primers. This is where whole genome sequencing (WGS) presents a huge advantage as it can identify bacteria as well as detect the presence of any AMR genes without prior knowledge 35 , and potentially without the need to culture.
Not all molecular techniques utilize nucleic acids as their form of detection, MALDI-TOF MS investigates the molecular composition of proteins and peptides within a sample. It identifies specific biomarkers based on their mass-to-charge ratio, providing information about the samples molecular profile 36 . This information used in conjunction with a reference database can be used to determine the identity of a pathogen and its AMR profile 37 . MALDI-TOF provides a comprehensive result with the potential to highlight multiple resistance mechanisms, however, this style of analysis can miss certain types of resistance that are not directly related to protein expression, such as mutations in regulatory regions or modifications in non-proteinaceous components of bacteria.
Sequencing and AMR prediction
Third generation WGS systems provide long reads at high speed, examples being Illumina MiniSeq & MiSeq, and Oxford Nanopore’s MiniON and PromethION 24 , 38 . These systems permit rapid pathogen identification and antibiotic resistance in a single assay without a culturing step. There is ever growing support in the AMR surveillance field that these WGS methods could replace current phenotypic assays 35 , 39 , 40 , 41 .
To identify which AMR genes are present post-sequencing requires two bioinformatic components: an aligner (e.g. Resistance Gene Identifier (RGI) 39 , AMRFinderPlus 42 and ResFinder 43 ) and a database of known AMR gene sequences and their associated resistance phenotype (e.g. Comprehensive Antibiotic Resistance Database (CARD) 39 , MegaRes 44 and National Database of Antibiotic Resistant Organisms (NDARO) 45 ). WGS analysis provides speed, flexibility, and breadth as clinical samples can be screened against hundreds of potential resistance profiles simultaneously – a process that would be too laborious and excessively time consuming for culture-based approaches. Furthermore, sequencing is neither dependent on pure cultures nor on being able to culture fastidious strains. WGS has great potential for AMR surveillance and diagnosis, but it is not a routine clinical application. Unlike direct phenotypic testing, sequencing predictions only indicate the presence (or absence) of antibiotic resistance sequences in the sample. Clinical and phenotypic information is usually required in order to properly interpret the outputs of sequencing. The presence of an AMR gene does not necessarily translate to antibiotic resistance since the genes may be inactive, an area where MALDI-TOF provides greater certainty of an active resistance owing to its detection of proteins that could be linked to a resistance genes expression 37 . Of greater concern is the observation that, the absence of any AMR indicator genes may not always correctly infer phenotypic susceptibility, a documented example of this can be found in the false negative predictions by WGS in Salmonella enterica 46 .
In 2017, EUCAST highlighted several issues that need to be addressed before the technology can move forward in the clinical context 47 . The key points were:
There is a lack of evidence for the AMR gene prediction accuracy for many bacteria.
It is a non-trivial process to establish the equivalent of clinical breakpoints in genomic predictions.
No standardisation of bioinformatics tools and approach to perform quality control (QC).
There is no single database of all known resistance genes/mutations - multiple databases developed independently means again that there is no harmonisation and data output is not equivalent.
Nevertheless, the cost of sequencing is coming down and the move towards high throughput methodologies is progressing, meaning that the current barriers to entry are reducing 48 . We are already seeing a strong push towards WGS/NGS sequencing in other diagnostic fields (e.g. genetic disorders) and the value it would add to AMR surveillance and evidence-based drug prescription is significant 49 , 50 . The creation of standards, both written and physical reference reagents, in this growing field would help to address many of the concerns of EUCAST and help to accelerate wider acceptance.
Real world examples of early AMR detection potential
To discuss how the different tools (Table 2 ) perform, we have chosen to highlight real world examples of where screening for antimicrobial resistance is essential or a growing concern:
Infection screening in blood and cerebrospinal fluid (CSF) samples
Blood and CSF samples are commonly used to diagnose bacterial infections, as healthy individuals harbour no bacteria from these sample sites. The normally sterile nature of CSF and blood in these sample sites makes them ideal samples for pathogenic strain detection and identification, as there is no bacterial background against which a pathogen needs to be distinguished from. Nevertheless, despite their diagnostic advantages, there often arises an urgent necessity to treat diseases associated with these samples, such as sepsis or numerous neonatal infections, due to their potentially life-threatening nature. Earlier and effective treatment results in better clinical outcomes, especially in younger patients who are at greater risk from bacterial infections 51 , 52 , 53 . It is standard practice for clinicians to begin empiric antibiotic treatment prior to receiving information on bacterial susceptibility.
A clinical example of CSF usage is Bacterial meningitis, a highly lethal disease 54 , 55 . Initial empirical treatment is often necessary with CSF samples taken prior to enable informed diagnosis. CSF culture is considered the gold standard; however, PCR is increasingly becoming relied upon because of its far greater sensitivity 55 . It is difficult to employ AMR stewardship, when delays in treatment can cause deaths, but we are beginning to see the results of this with third-generation antibiotics (e.g. ceftriaxone) becoming ineffective against Escherichia coli meningitis 56 . The high mortality rate associated with these infections means it is essential we try to move towards faster diagnostic tools to provide early, effective treatment based on evidence (reliable clinical laboratory test results). An ONT (Oxford Nanopore Technologies)-based approach to WGS and rapid diagnostics in blood infections is considered very promising with high accuracy and fast turnaround results, with the potential to be applied to CSF and implemented in clinical settings 57 .
Sexually transmitted infections (STIs)
STIs are a global problem, with the highest burden in low- and middle- income countries (LMICs). When left untreated, STIs can cause complications ranging from problems with fertility and pregnancy to cancer 58 . The most common bacterial STIs are Chlamydia trachomatis and Neisseria gonorrhoeae (inferred from Public Health England data 59 ). Treponema pallidum (syphilis) , Haemophilus ducreyi (Chancroid), and Mycoplasma genitalium infections are also prevalent but occur at a far lower frequency 59 .
The relatively few causative bacterial agents associated with STIs makes targeted NAAT-based diagnostics an effective solution for infection identification. Furthermore, the characterisation of common AMR causing genes found in C. trachomatis and N. gonorrhoeae , also lend themselves well to NAAT-based AMR detection 24 , 60 , 61 , 62 and the preferred laboratory method for these two strains has shifted from culture 63 to NAAT 64 increasing sensitivity and specificity, and faster turnaround time 63 . WGS offers an alternative that would be able to strain ID and screen for AMR at the same time (and rapidly), but the high incidence would be too expensive in comparison to NAAT. However, given the rise in novel AMR causing genes it may become necessary in the future. Indeed, “super” gonorrhoea is already a growing AMR concern, with the first case of drug resistant gonorrhoea reported in London in December 2021 65 and a further two cases in the UK as of the 7 th February 2022 and increasing numbers across Europe 66 . The World Health Organisation (WHO) has launched a global action plan to control the spread and impact of antimicrobial resistance in N. gonorrhoeae as part of a wider STI surveillance plan, with a focus on controlling antibiotic usage and disease spread.
Urinary tract infections (UTIs)
UTI infections are the most common infectious disease after respiratory tract infections and are a major public health problem in terms of morbidity and financial cost 67 . There has been an alarming rise in UTI antimicrobial resistance, likely owing to UTI patients being among the top receivers of outpatient antibiotic prescriptions 67 , 68 . The leading cause of UTI’s is uropathogenic Escherichia coli (UPEC), making up 80% of infections in women aged 18–39 69 , 70 . The current leading approach to identification and antibiotic susceptibility is culture-based screening. Given the overwhelming amount of UPEC caused infections and the small bacterial background of the sample one could argue for the use of lateral flow or multiplex PCR to confirm presence of E. coli and its resistance profile. Although neither of these approaches would rule out other organisms, they provide a far more rapid diagnosis of the leading cause. There is a growth of emerging technologies in UTI diagnostics, utilising microfluidics and lab-on-a-chip concepts to help provide point of care species identification and treatment suggestions 71 , 72 .
Upper and lower respiratory tract infections
Respiratory tract infections are a major global health issue, especially in low-income countries with limited healthcare access. Additionally, outbreaks of highly contagious respiratory infections, can have far-reaching consequences on a global scale, causing widespread illness, economic disruption, and loss of life. Efforts to prevent, detect, and effectively manage respiratory tract infections are crucial for safeguarding public health and minimising their impact. Pathogens commonly causing respiratory infections include Streptococcus pneumoniae , Haemophilus influenzae , Mycoplasma pneumoniae , Pseudomonas aeruginosa , and Mycobacterium tuberculosis 73 . Antibiotics are used to manage these infections with over half of all UK oral antibiotic prescriptions being written for this indication 74 . Several mechanisms conferring antimicrobial resistance in the organisms listed above have been observed and are of increasing concern 73 , 74 . A recent study describes the life cycle of antibiotic resistance genes in Pseudomonas aeruginosa isolated from hospitalised, ventilated patients 75 . They demonstrate the value of using targeted sequencing to identify and track AMR genes, showing that the data generated can inform treatment by enabling patient-specific antibiotic cycling strategies.
Pulmonary tuberculosis (TB) can be transmitted between individuals and is a significant contributor to poor health and mortality rates globally. Prior to the COVID-19 pandemic, TB held the unfortunate distinction of being the leading cause of death among single agent infectious diseases, surpassing even HIV/AIDS in its impact 76 . Culture of sputum of other respiratory secretions is considered the gold standard for diagnosis, however, it is slow-growing, taking two to six weeks for culture and an additional three plus weeks for multi-drug resistance testing 77 . Rapid detection of resistance patterns and prompt initiation of appropriate treatment are essential for effectively controlling TB and minimising the transmission of drug-resistant strains 78 . Faster diagnostic and susceptibility assays already exist (both NAAT and WGS based 79 ), with the WHO now pushing for better access to rapid testing 80 . The UK is leading on that front, having implemented the first service for TB rapid diagnostics utilising WGS, shortening TB diagnosis and treatment in a cost-effective way 81 . This successful implementation is a promising first step for a wider adoption of rapid diagnostics in healthcare for other indications.
Skin and soft tissue infections
Conditions affecting the skin can be both physically painful and disfiguring, leading to both physical discomfort, mental distress and social isolation. Among medical practitioners, dermatologists have the highest prescription rates for antibiotics 82 . Nosocomial (healthcare-associated) infections are a serious complication of severe burns, and the use of systemic prophylactic antibiotics to control infections and reduce sepsis risk has been discussed in several studies over the past several years 83 . These studies have shown that prophylactic use of broad-spectrum antibiotics does not provide protection against sepsis, except for patients with inhalation burns or pneumonia. The overarching theme here is that broad spectrum antibiotics are often used for skin conditions, with little diagnostic or susceptibility screening performed. This needs to change to prevent further AMR evolution. Targeted therapy based on identification of the causative agent and its susceptibility will need to increase in importance.
Chronic liver disease
Bacterial infections are common in patients with chronic liver disease (CLD) and are one of the most important causes of liver-related complications, progression to liver failure, and mortality in these patients 84 . Resorting to antibiotic prophylaxis and broad-spectrum empirical therapy remains essential in the management of infection prevention in advanced CLD 85 . This approach has however led to the widespread use of antimicrobials, which is the leading cause of the continuous rise of MDRO infections. MDRO rates to quinolone drugs have been recorded up to 40% in CLD patients with spontaneous bacterial peritonitis on prophylactic antibiotics, leading to a break-through recurrence of intra-peritoneal infection. MDR bacteria have emerged as a significant challenge in many countries 86 , and infections caused by these bacteria are associated with a particularly poor prognosis in CLD patients 87 .
Circumventing the harmful impact of AMR in CLD requires a combinatorial approach encompassing antibiotic stewardship programmes, accurate biomarkers of infection onset and resolution, prompting the rapid de-escalation of antimicrobial therapies 88 , 89 . The other crucial aspect remains the development of rapid testing technologies for the accurate identification of causative pathogens with simultaneous AMR profiling to guide timely and accurate antibacterial therapy in cirrhosis patients 90 This has paved the way for non-culture-based approaches that offer the potential in reducing the limitations, delays and inaccuracies that are associated with conventional microbiological techniques 91 .
Microbiome donor screening
Microbiome therapies are a growing area in medicine, offering novel approaches to disease management in instances of unmet clinical need or poor treatment outcomes 92 . Faecal microbiota transplant (FMT) is the first commercially available microbiome treatment, employed to treat recurrent Clostridioides difficile infections 93 . In 2019, the first death caused by FMT occurred in the USA when an immunocompromised adult received a FMT that led to an invasive infection caused by extended-spectrum beta-lactamase (ESBL)-producing E. coli present in the donor stool 94 . This triggered the FDA to recommend screening for common MDROs 95 in all donor samples, with the British Society of Gastroenterology and Healthcare Infection Society adopting similar recommendations 96 , 97 . Even with all these guidelines and safety measures now in place, cases of Shiga toxin E. coli infections caused by FMT are still occurring 98 , 99 , raising concern that pre-treatment screening is not sufficiently robust. Culture based testing for many bacteria may not be consistent or reliable 100 , 101 . In the case of FMT, false negatives become an unacceptable risk, with the danger of transmitting an infectious organism that was not detected during donor screening.
Molecular methods should be adopted for FMT screening (both for identification of pathogens and AMR assessment) since the sensitivity of this method is much higher than culture. WGS could add further value as it not only identifies strains in a complex mixture and screens for markers of AMR, but it can also provide data that could be used to identify microbiome dysbiosis, potentially before a disease has manifested. Other treatments using microbiome transplants (e.g., vaginal microbiome transplants for bacterial vaginosis) would also benefit from the availability of WGS screening and characterisation (strain ID and AMR).
Standardisation
The need for standards in antibiotic resistance gene detection by WGS is crucial in combating the global threat of antimicrobial resistance. WGS has emerged as a powerful tool for identifying and characterising antibiotic resistance genes in bacterial genomes. However, the lack of standardised protocols and guidelines for WGS-based resistance profile detection is hindering the accurate and consistent interpretation of results. Biological standardisation is necessary to ensure that different laboratories and researchers are harmonised in quality control measures and data analysis pipelines 102 , 103 . This will enable reliable comparisons of resistance profiles across studies and facilitate the development of robust surveillance systems. Furthermore, standardised protocols and the use of appropriate reference reagents will promote data sharing and collaboration, allowing for the accumulation of comprehensive and representative datasets that can inform evidence-based policies and interventions. Ultimately, the establishment of written and physical standards in antibiotic resistance gene detection by WGS will enhance our understanding of the global antimicrobial resistance landscape and support efforts to mitigate its impact on public health.
The rise of multi-drug resistant organisms (MDROs) poses a significant threat to global health, leading to increased mortality rates, healthcare and societal costs which necessitate radical intervention. Current methods for AMR detection, most significantly culture-based approaches, have limitations in terms of sensitivity, turnaround time, and the ability to detect all potential resistance genes. WGS offers a promising alternative, providing rapid and comprehensive information about the presence of AMR genes in bacterial strains (Fig. 1 ). However, several challenges need to be addressed before WGS can be widely implemented in clinical settings. These include the need for standardised methodologies, a comprehensive and unified database of known resistance genes, the availability of appropriate physical reference materials to assure assay performance and the establishment of clinical breakpoints for genomic predictions. Additionally, the cost of sequencing and the interpretation of sequencing results need to be considered to ensure LMICs can also access and derive maximal benefit, as well as optimising upstream processes including biological sample handling and DNA extraction. Despite these challenges, the potential benefits of WGS in AMR surveillance and evidence-based antimicrobial prescription are significant. Establishing standards for WGS-based AMR detection will help address these challenges and accelerate the adoption of this powerful tool in the fight against AMR, ultimately leading to more effective and targeted treatment strategies.
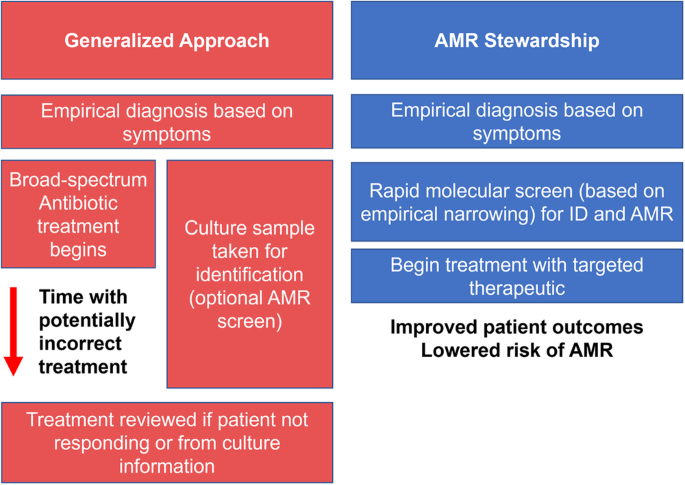
Current approach for infection diagnostics vs a WGS approach that supports AMR stewardship.
Ventola, C. L. The antibiotic resistance crisis: causes and threats. P & T J. (2015).
Sengupta, S., Chattopadhyay, M. K. & Grossart, H. P. The multifaceted roles of antibiotics and antibiotic resistance in nature. Front. Microbiol. Preprint at https://doi.org/10.3389/fmicb.2013.00047 (2013).
Michael, C. A., Dominey-Howes, D. & Labbate, M. The antimicrobial resistance crisis: causes, consequences, and management. Front Public Health 2 , 145 (2014).
Article PubMed PubMed Central Google Scholar
Magiorakos, A. P. et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Inf. https://doi.org/10.1111/j.1469-0691.2011.03570.x (2012).
Laxminarayan, R. et al. The Lancet Infectious Diseases Commission on antimicrobial resistance: 6 years later. Lancet Inf. Dis. Preprint at https://doi.org/10.1016/S1473-3099(20)30003-7 (2020).
Roca, I. et al. The global threat of antimicrobial resistance: science for intervention. N. Microbes N. Infect. 6 , 22–29 (2015).
Article CAS Google Scholar
Pokharel, S., Shrestha, P. & Adhikari, B. Antimicrobial use in food animals and human health: time to implement ‘One Health’ approach. Antimicrob. Resist Infect. Control 9 , 181 (2020).
Munkholm, L. & Rubin, O. The global governance of antimicrobial resistance: a cross-country study of alignment between the global action plan and national action plans. Global Health https://doi.org/10.1186/s12992-020-00639-3 (2020).
Seventy-first UN General Assembly. Political declaration of the high-level meeting of the General Assembly on antimicrobial resistance . United Nations. Preprint at (2016).
Robinson, T. P. et al. Antibiotic resistance is the quintessential One Health issue. Trans. R. Soc. Trop. Med. Hyg. Preprint at https://doi.org/10.1093/trstmh/trw048 (2016).
Velazquez-Meza, M. E., Galarde-López, M., Carrillo-Quiróz, B. & Alpuche-Aranda, C. M. Antimicrobial resistance: One Health approach. Vet. World 15 , 743–749 (2022).
Article CAS PubMed PubMed Central Google Scholar
Naylor, N. R. et al. Estimating the burden of antimicrobial resistance: a systematic literature review. Antimicrob. Resist Infect Control https://doi.org/10.1186/s13756-018-0336-y (2018).
IACG. No Time To Wait: Infections From Drug-Resistant Securing the Future From Drug-Resistant Infections. Artforum International (2019).
Hendriksen, R. S. et al. Global monitoring of antimicrobial resistance based on metagenomics analyses of urban sewage. Nat. Commun. 10 , 1124 (2019).
Burnham, J. P., Olsen, M. A. & Kollef, M. H. Re-estimating annual deaths due to multidrug-resistant organism infections. Infect. Control Hosp. Epidemiol. 40 , 112–113 (2019).
Article PubMed Google Scholar
Folgori, L. et al. Healthcare-associated infections in pediatric and neonatal intensive care units: impact of underlying risk factors and antimicrobial resistance on 30-day case-fatality in italy and brazil. Infect Control Hosp Epidemiol https://doi.org/10.1017/ice.2016.185 (2016).
Yusef, D. et al. Community-acquired serious bacterial infections in the first 90 days of life: a revisit in the era of multi-drug-resistant organisms. World J. Ped. https://doi.org/10.1007/s12519-019-00276-w (2019).
Qureshi, S., Maria, N., Zeeshan, M., Irfan, S. & Qamar, F. N. Prevalence and risk factors associated with multi-drug resistant organisms (MDRO) carriage among pediatric patients at the time of admission in a tertiary care hospital of a developing country. A cross-sectional study. BMC Infect. Dis. https://doi.org/10.1186/s12879-021-06275-5 (2021).
Nations, U., of Economic, D., Affairs, S. & Division, P. World Urbanization Prospects The 2018 Revision. (2018).
Smith, K. F. et al. Global rise in human infectious disease outbreaks. J. R. Soc. Interface 11 , 20140950 (2014).
Peters, A., Vetter, P., Guitart, C., Lotfinejad, N. & Pittet, D. Understanding the emerging coronavirus: what it means for health security and infection prevention. J. Hosp. Infect. 104 , 440–448 (2020).
Ventola, C. L. The antibiotic resistance crisis: part 1: causes and threats. P T 40 , 277–283 (2015).
PubMed PubMed Central Google Scholar
Giske, C. G. et al. EUCAST guidelines for detection of resistance mechanisms and specific resistances of clinical and/or epidemiological importance. The European Committee on Antimicrobial Susceptibility Testing (2017).
Vasala, A., Hytönen, V. P. & Laitinen, O. H. Modern Tools for Rapid Diagnostics of Antimicrobial Resistance. Front. Cellular Infect. Microbiol. Preprint at https://doi.org/10.3389/fcimb.2020.00308 (2020).
CLSI. Methods for Antimicrobial Susceptibility Testing of Anaerobic Bacteria; Approved Standard - Eighth Edition . CLSI Document M11-A8 (2012).
Knight, G. M. et al. Fast and expensive (PCR) or cheap and slow (culture)? A mathematical modelling study to explore screening for carbapenem resistance in UK hospitals. BMC Med. https://doi.org/10.1186/s12916-018-1117-4 (2018).
Vasala, A., Hytönen, V. P. & Laitinen, O. H. Modern Tools for Rapid Diagnostics of Antimicrobial Resistance. Front Cell Infect Microbiol . 10 (2020).
Franco-Duarte, R. et al. Advances in chemical and biological methods to identify microorganisms—from past to present. Microorganisms 7 , 130 (2019).
Lee, J. Y. et al. Diagnostic yield of stool culture and predictive factors for positive culture in patients with diarrheal illness. In Medicine (United States) https://doi.org/10.1097/MD.0000000000007641 (2017).
Hsiao, W. W. W. et al. Recent advances in novel lateral flow technologies for detection of COVID-19. Biosensors. Preprint at https://doi.org/10.3390/bios11090295 (2021).
Andryukov, B. G. Six decades of lateral flow immunoassay: From determining metabolic markers to diagnosing covid-19. AIMS Microbiol. Preprint at https://doi.org/10.3934/microbiol.2020018 (2020).
Boutal, H. et al. Development and validation of a lateral flow immunoassay for rapid detection of NDM-producing enterobacteriaceae. J. Clin. Microbiol. https://doi.org/10.1128/JCM.00248-17 (2017).
Yang, S. & Rothman, R. E. PCR-based diagnostics for infectious diseases: Uses, limitations, and future applications in acute-care settings. Lancet Infect. Dis. Preprint at https://doi.org/10.1016/S1473-3099(04)01044-8 (2004).
Chan, K. et al. A rapid and low-cost PCR thermal cycler for infectious disease diagnostics. PLoS One https://doi.org/10.1371/journal.pone.0149150 (2016).
Hendriksen, R. S. et al. Using Genomics to Track Global Antimicrobial Resistance. Front. Public Health 7 , 242 (2019).
Wieser, A., Schneider, L., Jung, J. & Schubert, S. MALDI-TOF MS in microbiological diagnostics—identification of microorganisms and beyond (mini review). Appl. Microbiol. Biotechnol. 93 , 965–974 (2012).
Article CAS PubMed Google Scholar
Yoon, E.-J. & Jeong, S. H. MALDI-TOF mass spectrometry technology as a tool for the rapid diagnosis of antimicrobial resistance in bacteria. Antibiotics 10 , 982 (2021).
van Belkum, A. & Rochas, O. Laboratory-based and point-of-care testing for MSSA/MRSA detection in the age of whole genome sequencing. Front. Microbiol. Preprint at https://doi.org/10.3389/fmicb.2018.01437 (2018).
McArthur, A. G. et al. The comprehensive antibiotic resistance database. Antimicrob. Agents Chemother. 57 , 3348–3357 (2013).
Alcock, B. P. et al. CARD 2020: antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res. 48 , D517–D525 (2019).
PubMed Central Google Scholar
Gupta, S. K. et al. ARG-ANNOT, a new bioinformatic tool to discover antibiotic resistance genes in bacterial genomes. Antimicrob. Agents Chemother. 58 , 212–220 (2014).
Feldgarden, M. et al. Curation of the AMRFinderPlus databases: applications, functionality and impact. Microb. Genom. 8 , (2022).
Florensa, A. F., Kaas, R. S., Clausen, P. T. L. C., Aytan-Aktug, D. & Aarestrup, F. M. ResFinder – an open online resource for identification of antimicrobial resistance genes in next-generation sequencing data and prediction of phenotypes from genotypes. Microb. Genom. 8 , (2022).
Bonin, N. et al. MEGARes and AMR++, v3.0: an updated comprehensive database of antimicrobial resistance determinants and an improved software pipeline for classification using high-throughput sequencing. Nucleic Acids Res. 51 , D744–D752 (2023).
Papp, M. & Solymosi, N. Review and comparison of antimicrobial resistance gene databases. Antibiotics 11 , 339 (2022).
Zwe, Y. H. et al. Whole genome sequencing (WGS) fails to detect antimicrobial resistance (AMR) from heteroresistant subpopulation of Salmonella enterica. Food Microbiol. 91 , 103530 (2020).
Ellington, M. J. et al. The role of whole genome sequencing in antimicrobial susceptibility testing of bacteria: report from the EUCAST Subcommittee. Clin. Microbiol. Infect. 23 , 2–22 (2017).
Li, H. et al. Cost-reduction strategies in massive genomics experiments. Mar. Life Sci. Technol. https://doi.org/10.1007/s42995-019-00013-2 (2019).
Pruneri, G. et al. Next-Generation Sequencing in Clinical Practice: Is It a Cost-Saving Alternative to a Single-Gene Testing Approach? Pharmacoecon. Open https://doi.org/10.1007/s41669-020-00249-0 (2021).
Xue, Y., Ankala, A., Wilcox, W. R. & Hegde, M. R. Solving the molecular diagnostic testing conundrum for Mendelian disorders in the era of next-generation sequencing: Single-gene, gene panel, or exome/genome sequencing. Gen. Med. https://doi.org/10.1038/gim.2014.122 (2015).
Edmond, K. & Zaidi, A. New approaches to preventing, diagnosing, and treating neonatal sepsis. PLoS Med . https://doi.org/10.1371/journal.pmed.1000213 (2010).
Darmstadt, G. L. et al. Evidence-based, cost-effective interventions: How many newborn babies can we save? Lancet https://doi.org/10.1016/S0140-6736(05)71088-6 (2005).
Simen-Kapeu, A. et al. Treatment of neonatal infections: A multi-country analysis of health system bottlenecks and potential solutions. BMC Pregnancy Childbirth https://doi.org/10.1186/1471-2393-15-S2-S6 (2015).
Okike, I. O. et al. Trends in bacterial, mycobacterial, and fungal meningitis in England and Wales 2004–11: an observational study. Lancet Infect. Dis. 14 , 301–307 (2014).
Griffiths, M. J., McGill, F. & Solomon, T. Management of acute meningitis. Clin. Med (Lond.) 18 , 164–169 (2018).
Liu, Y. et al. Escherichia coli causing neonatal meningitis during 2001-2020: a study in Eastern China. Int J. Gen. Med 14 , 3007–3016 (2021).
Taxt, A. M., Avershina, E., Frye, S. A., Naseer, U. & Ahmad, R. Rapid identification of pathogens, antibiotic resistance genes and plasmids in blood cultures by nanopore sequencing. Sci. Rep. 10 , 7622 (2020).
Chesson, H. W., Mayaud, P. & Aral, S. O. Sexually transmitted infections: impact and cost-effectiveness of prevention. in Disease Control Priorities, Third Edition (Volume 6): Major Infectious Diseases 203–232 (The World Bank, 2017). https://doi.org/10.1596/978-1-4648-0524-0_ch10 .
Public Health England. STI Diagnoses and Rates in England by Gender, 2011 to 2020 . (2021).
Hemarajata, P., Yang, S., Soge, O. O., Humphries, R. M. & Klausner, J. D. Performance and Verification of a Real-Time PCR Assay Targeting the gyrA Gene for Prediction of Ciprofloxacin Resistance in Neisseria gonorrhoeae. J. Clin. Microbiol. https://doi.org/10.1128/JCM.03032-15 (2016).
Donà, V. et al. Multiplex real-time PCR assay with high-resolution melting analysis for characterization of antimicrobial resistance in neisseria gonorrhoeae. J. Clin. Microbiol. https://doi.org/10.1128/JCM.03354-15 (2016).
Benamri, I., Azzouzi, M., Sanak, K., Moussa, A. & Radouani, F. An overview of genes and mutations associated with Chlamydiae species’ resistance to antibiotics. Annals Clin. Microbiol. Antimicrob. https://doi.org/10.1186/s12941-021-00465-4 (2021).
Ng, L. K. & Martin, I. E. The laboratory diagnosis of Neisseria gonorrhoeae. Can. J. Infect. Dis. Med. Microbiol. 16 , 15 (2005).
Papp, J. R., Schachter, J., Gaydos, C. A. & Van Der Pol, B. Recommendations for the Laboratory-Based Detection of Chlamydia trachomatis and Neisseria gonorrhoeae-2014. MMWR Recomm. Rep. 63 , 1–19 (2014).
Google Scholar
UKHSA UK Health Security Agency. Antibiotic-resistant strain of gonorrhoea detected in London - GOV.UK. Press Release https://www.gov.uk/government/news/antibiotic-resistant-strain-of-gonorrhoea-detected-in-london?fbclid=IwAR0M_SJ8KosA-MEm3ayjjpeuO8Go9xgyq1ExjN_VGXW3Ak1APfOL_4-yARc (2021).
UK Health Security Agency. More cases of antibiotic resistant gonorrhoea identified in England. Press release (Gov.uk) https://www.gov.uk/government/news/more-cases-of-antibiotic-resistant-gonorrhoea-identified-in-england (2022).
Toosky, M. N. et al. A rapid, point-of-care antibiotic susceptibility test for urinary tract infections. J. Med. Microbiol. https://doi.org/10.1099/jmm.0.001119 (2020).
Grigoryan, L., Trautner, B. W. & Gupta, K. Diagnosis and management of urinary tract infections in the outpatient setting: A review. JAMA-J. Am. Med. Assoc. https://doi.org/10.1001/jama.2014.12842 (2014).
Stamm, W. E. Scientific and clinical challenges in the management of urinary tract infections. Am. J. Med. https://doi.org/10.1016/s0002-9343(02)01053-7 (2002).
Foxman, B. The epidemiology of urinary tract infection. Nat. Rev. Urol. https://doi.org/10.1038/nrurol.2010.190 (2010).
Davenport, M. et al. New and developing diagnostic technologies for urinary tract infections. Nat. Rev. Urol. 14 , 296–310 (2017).
Santos, M. et al. A review on urinary tract infections diagnostic methods: Laboratory-based and point-of-care approaches. J. Pharm. Biomed. Anal . 219 https://doi.org/10.1016/j.jpba.2022.114889 (2022).
Guitor, A. K. & Wright, G. D. Antimicrobial Resistance and Respiratory Infections. Chest 154 , 1202–1212 (2018).
Derbyshire, E. J. & Calder, P. C. Respiratory Tract Infections and Antibiotic Resistance: A Protective Role for Vitamin D? Front. Nutr. 8 , (2021).
Chung, H. et al. Rapid expansion and extinction of antibiotic resistance mutations during treatment of acute bacterial respiratory infections. Nat. Commun. 13 , 1231 (2022).
World Health Organization. Global Tuberculosis Report 2022 . https://www.who.int/publications/i/item/9789240061729 (2022).
CADTH. Rapid and Simultaneous Tuberculosis and Antibiotic Susceptibility Testing for the Diagnosis of Pulmonary Tuberculosis and Rifampicin Resistance: A Review of Diagnostic Accuracy . (2020).
Finci, I. et al. Investigating resistance in clinical Mycobacterium tuberculosis complex isolates with genomic and phenotypic antimicrobial susceptibility testing: a multicentre observational study. Lancet Microbe 3 , e672–e682 (2022).
Zhao, K. et al. Rapid identification of drug-resistant tuberculosis genes using direct PCR amplification and oxford nanopore technology sequencing. Can. J. Infect. Dis. Med. Microbiol. 2022 , 7588033 (2022).
World Health Organisation (WHO). WHO Operational Handbook on Tuberculosis Module 3: Diagnosis . (WHO, 2021).
Mugwagwa, T., Abubakar, I. & White, P. J. Using molecular testing and whole-genome sequencing for tuberculosis diagnosis in a low-burden setting: a cost-effectiveness analysis using transmission-dynamic modelling. Thorax 76 , 281–291 (2021).
Muhaj, F. F., George, S. J. & Tyring, S. K. Bacterial antimicrobial resistance and dermatological ramifications*. Br. J. Dermatol. 187 , 12–20 (2022).
Muthukumar, V., Arumugam, P. K. & Bamal, R. Role of systemic antibiotic prophylaxis in acute burns: A retrospective analysis from a tertiary care center. Burns 46 , 1060–1065 (2020).
Jalan, R. et al. Bacterial infections in cirrhosis: A position statement based on the EASL Special Conference 2013. J. Hepatol. 60 https://doi.org/10.1016/j.jhep.2014.01.024 (2014).
Patel, V. C. et al. Rifaximin-α reduces gut-derived inflammation and mucin degradation in cirrhosis and encephalopathy: RIFSYS randomised controlled trial. J. Hepatol. 76 , (2022).
Acevedo, J. Multiresistant bacterial infections in liver cirrhosis: Clinical impact and new empirical antibiotic treatment policies. World J. Hepatol. 7 , (2015).
Salerno, F. et al. The impact of infection by multidrug-resistant agents in patients with cirrhosis. A multicenter prospective study. Liver International 37 , (2017).
Bajaj, J. S., Kamath, P. S. & Reddy, K. R. The Evolving Challenge of Infections in Cirrhosis. New Engl. J. Med. 384 , (2021).
Patel, V. C. & Williams, R. Antimicrobial resistance in chronic liver disease. Hepatol. Int. 14 https://doi.org/10.1007/s12072-019-10004-1 (2020).
Shallcross, L. & O’Brien, A. Antimicrobial resistance in liver disease: better diagnostics are needed. Lancet Gastroenterol. Hepatol. 2 https://doi.org/10.1016/S2468-1253(16)30240-0 (2017).
van Belkum, A. et al. Developmental roadmap for antimicrobial susceptibility testing systems. Nat. Rev. Microbiol. 17 , (2019).
Surawicz, C., Bowman, K. & Broussard, E. Fecal microbiota transplantation: current clinical efficacy and future prospects. Clin. Exp. Gastroenterol. 8 , 285–291 (2015).
Pérez-Cobas, A., Moya, A., Gosalbes, M. & Latorre, A. Colonization Resistance of the Gut Microbiota against Clostridium difficile. Antibiotics 4 , 337–357 (2015).
FDA. Important safety alert regarding use of fecal microbiota for transplantation and risk of serious adverse reactions due to transmission of multi-drug resistant organisms | FDA. FDA Safety & Availability (Biologics) (2019).
FDA. Information Pertaining to Additional Safety Protections Regarding Use of Fecal Microbiota for Transplantation – Screening and Testing of Stool Donors for Multi-drug Resistant Organisms. FDA Safety & Availability (Biologics) (2019).
Mullish, B. H. et al. The use of faecal microbiota transplant as treatment for recurrent or refractory Clostridium difficile infection and other potential indications: joint British Society of Gastroenterology (BSG) and Healthcare Infection Society (HIS) guidelines. Gut 67 , 1920–1941 (2018).
Cammarota, G. et al. International consensus conference on stool banking for faecal microbiota transplantation in clinical practice. Gut 68 , 2111–2121 (2019).
Hohmann, E. L. Faecal microbiota transplantation: more screening for old and new pathogens. Lancet Inf. Dis. https://doi.org/10.1016/S1473-3099(20)30850-1 (2021).
FDA. Fecal Microbiota for Transplantation: Safety Alert - Risk of Serious Adverse Events Likely Due to Transmission of Pathogenic Organisms. FDA Safety & Availability (Biologics) (2020).
Arjyal, A. et al. Gatifloxacin versus chloramphenicol for uncomplicated enteric fever: An open-label, randomised, controlled trial. Lancet Infect Dis . https://doi.org/10.1016/S1473-3099(11)70089-5 (2011).
World Health Organization. The Selection and Use of Essential Medicines . WHO Technical Report Series 1021 (2020).
Amos, G. C. A. et al. Developing standards for the microbiome field. Microbiome https://doi.org/10.1186/s40168-020-00856-3 (2020).
Sergaki, C. et al. Developing whole cell standards for the microbiome field. Microbiome 10 , 1–16 (2022).
Article Google Scholar
Download references
Acknowledgements
We would like to acknowledge Dr Neil Almond for the support and the feedback on the document.
Author information
Authors and affiliations.
Science Research and Innovation, Medicines and Healthcare products Regulatory Agency, Blanche Lane, South Mimms, Potters Bar, Hertfordshire, EN6 3QG, UK
Jack Hassall, Carmen Coxon & Chrysi Sergaki
The Roger Williams Institute of Hepatology London, Foundation for Liver Research, 111 Coldharbour Lane, London, SE5 9NT, UK
Vishal C. Patel
Institute of Liver Studies, School of Immunology and Microbial Sciences, Faculty of Life Sciences and Medicine, King’s College London, 125 Coldharbour Lane, London, SE5 9NU, UK
Institute of Liver Studies, King’s College Hospital NHS Foundation Trust, Denmark Hill, London, SE5 9RS, UK
Centre for Clinical Infection and Diagnostics Research, Guy’s and St Thomas’ NHS Foundation Trust and King’s College, London, UK
Simon D. Goldenberg
You can also search for this author in PubMed Google Scholar
Contributions
C.S. and J.H. conceived the idea and formatted the structure of the manuscript. JH wrote the main body, C.C., V.P. and S.G. reviewed and added to the manuscript. C.S. reviewed and refined the manuscript. The authors read and approved the final manuscript.
Corresponding author
Correspondence to Chrysi Sergaki .
Ethics declarations
Competing interests.
The Medicines and Healthcare product Regulatory Authority laboratories at South Mimms is part of an Arms Length Body of the UK Government. The laboratories prepare, curate and distribute globally, reference materials including WHO International Standards. The distribution of reference materials is undertaken on a cost-recovery basis. Further information is available at www.nibsc.org All authors declare no financial or non-financial competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/ .
Reprints and permissions
About this article
Cite this article.
Hassall, J., Coxon, C., Patel, V.C. et al. Limitations of current techniques in clinical antimicrobial resistance diagnosis: examples and future prospects. npj Antimicrob Resist 2 , 16 (2024). https://doi.org/10.1038/s44259-024-00033-8
Download citation
Received : 20 November 2023
Accepted : 07 May 2024
Published : 17 June 2024
DOI : https://doi.org/10.1038/s44259-024-00033-8
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Sign up for the Nature Briefing: Microbiology newsletter — what matters in microbiology research, free to your inbox weekly.

- eLibrary Home
- eLibrary Login
- < Previous Event
- Next Event >
Home > Conferences > AMCIS > AMCIS 2024 Proceedings > IS in Educ, IS Curriculum, and Teaching Cases (SIG ED) > 12

IS in Educ, IS Curriculum, and Teaching Cases (SIG ED)
Proposal - Security Certifications, Degrees, & Work Experience: Which is better?
Presenter Information
Garry White , Texas State Univeristy Follow
Description
In this document we describe the formatting requirements for the Proceedings of the AMCIS Conference. Please review this document carefully. You can use this document as a template and copy/paste your paper content here (this might be the best or easiest way). Please be sure to adhere to the formatting requirements as this will be your camera-ready version, and a PDF format will be generated directly from your submitted final Word version. Please note several limitations on length: (1) your abstract should be no more than 150 words, as the abstract will also be used for the conference program, (2) your completed research paper should be no more than 10 pages (approx. 5,000 words, including figures, tables, references, and appendices). Emergent Research Forum papers are limited to 5 pages (approx. 2,500 words, including figures, tables, references, and appendices.) This paper length is intended to encourage authors to publish full-length papers in journals or other outlets later.
Paper Number
Recommended citation.
White, Garry, "Proposal - Security Certifications, Degrees, & Work Experience: Which is better?" (2024). AMCIS 2024 Proceedings . 12. https://aisel.aisnet.org/amcis2024/is_education/is_education/12
Since June 13, 2024
- https://amcis2024.aisconferences.org/
Advanced Search
- Notify me via email or RSS
- All Content
Author Corner
- eLibrary FAQ
- AMCIS 2024 Conference
Home | About | FAQ | My Account | Accessibility Statement
Privacy Copyright
- SUGGESTED TOPICS
- The Magazine
- Newsletters
- Managing Yourself
- Managing Teams
- Work-life Balance
- The Big Idea
- Data & Visuals
- Reading Lists
- Case Selections
- HBR Learning
- Topic Feeds
- Account Settings
- Email Preferences
Build a Corporate Culture That Works

There’s a widespread understanding that managing corporate culture is key to business success. Yet few companies articulate their culture in such a way that the words become an organizational reality that molds employee behavior as intended.
All too often a culture is described as a set of anodyne norms, principles, or values, which do not offer decision-makers guidance on how to make difficult choices when faced with conflicting but equally defensible courses of action.
The trick to making a desired culture come alive is to debate and articulate it using dilemmas. If you identify the tough dilemmas your employees routinely face and clearly state how they should be resolved—“In this company, when we come across this dilemma, we turn left”—then your desired culture will take root and influence the behavior of the team.
To develop a culture that works, follow six rules: Ground your culture in the dilemmas you are likely to confront, dilemma-test your values, communicate your values in colorful terms, hire people who fit, let culture drive strategy, and know when to pull back from a value statement.
Start by thinking about the dilemmas your people will face.
Idea in Brief
The problem.
There’s a widespread understanding that managing corporate culture is key to business success. Yet few companies articulate their corporate culture in such a way that the words become an organizational reality that molds employee behavior as intended.
What Usually Happens
How to fix it.
Follow six rules: Ground your culture in the dilemmas you are likely to confront, dilemma-test your values, communicate your values in colorful terms, hire people who fit, let culture drive strategy, and know when to pull back from a value.
At the beginning of my career, I worked for the health-care-software specialist HBOC. One day, a woman from human resources came into the cafeteria with a roll of tape and began sticking posters on the walls. They proclaimed in royal blue the company’s values: “Transparency, Respect, Integrity, Honesty.” The next day we received wallet-sized plastic cards with the same words and were asked to memorize them so that we could incorporate them into our actions. The following year, when management was indicted on 17 counts of conspiracy and fraud, we learned what the company’s values really were.
- EM Erin Meyer is a professor at INSEAD, where she directs the executive education program Leading Across Borders and Cultures. She is the author of The Culture Map: Breaking Through the Invisible Boundaries of Global Business (PublicAffairs, 2014) and coauthor (with Reed Hastings) of No Rules Rules: Netflix and the Culture of Reinvention (Penguin, 2020). ErinMeyerINSEAD
Partner Center
Information
- Author Services
Initiatives
You are accessing a machine-readable page. In order to be human-readable, please install an RSS reader.
All articles published by MDPI are made immediately available worldwide under an open access license. No special permission is required to reuse all or part of the article published by MDPI, including figures and tables. For articles published under an open access Creative Common CC BY license, any part of the article may be reused without permission provided that the original article is clearly cited. For more information, please refer to https://www.mdpi.com/openaccess .
Feature papers represent the most advanced research with significant potential for high impact in the field. A Feature Paper should be a substantial original Article that involves several techniques or approaches, provides an outlook for future research directions and describes possible research applications.
Feature papers are submitted upon individual invitation or recommendation by the scientific editors and must receive positive feedback from the reviewers.
Editor’s Choice articles are based on recommendations by the scientific editors of MDPI journals from around the world. Editors select a small number of articles recently published in the journal that they believe will be particularly interesting to readers, or important in the respective research area. The aim is to provide a snapshot of some of the most exciting work published in the various research areas of the journal.
Original Submission Date Received: .
- Active Journals
- Find a Journal
- Proceedings Series
- For Authors
- For Reviewers
- For Editors
- For Librarians
- For Publishers
- For Societies
- For Conference Organizers
- Open Access Policy
- Institutional Open Access Program
- Special Issues Guidelines
- Editorial Process
- Research and Publication Ethics
- Article Processing Charges
- Testimonials
- Preprints.org
- SciProfiles
- Encyclopedia

Article Menu

- Subscribe SciFeed
- Recommended Articles
- Google Scholar
- on Google Scholar
- Table of Contents
Find support for a specific problem in the support section of our website.
Please let us know what you think of our products and services.
Visit our dedicated information section to learn more about MDPI.
JSmol Viewer
Modeling and research on offshore casing cutting of hydraulic internal cutting device.

1. Introduction
2. mechanical casing cutting device, 2.1. basic structure of mechanical casing cutting device, 2.2. working principle, 3. the theory model of casing cutting, 3.1. the relationship between piston displacement and cutting tool tip radius, 3.2. calculation of cutting torque, 3.3. calculation of wellhead driving torque, 4. the 2d cutting simulation based on abaqus, 4.1. theoretical model of cutting, 4.2. simulation model and boundary conditions based on abaqus, 4.3. simulation analysis and results based on abaqus, 4.3.1. the influence of different tool rotational speeds on cutting simulation, 4.3.2. the impact of different cutting depths on cutting simulation, 4.3.3. the impact of different tool front angles on cutting simulation, 4.3.4. summary of the chapter, 5. analysis of influencing factors on the cutting efficiency of the cutting tool, 5.1. the cutter face angle α, 5.2. the driving force of drilling fluid f 0, 5.3. the cutting depth l and revolution of drill string n, 6. case study and discussion, 6.1. field casing cutting operation condition, 6.2. torque comparison at different rotational speeds, 7. conclusions, author contributions, informed consent statement, data availability statement, conflicts of interest.
- John, P. Health and safety at work offshore in Greg Gordon. In Oil and Gas law: Current Practice and Emerging Trends ; John, P., Emreusenmez, Eds.; Dundee University Press: Dundee, UK, 2011; pp. 193–219. [ Google Scholar ]
- Qiu, R.; Li, Z.; Zhang, Q.; Yao, X.; Xie, S.; Liao, Q.; Wang, B. A Realistic and Integrated Model for Evaluating Offshore Oil Development. J. Mar. Sci. Eng. 2022 , 10 , 1155. [ Google Scholar ] [ CrossRef ]
- Akande, K.O.; Owolabi, T.O.; Olatunji, S.O.; AbdulRaheem, A. A hybrid particle swarm optimization and support vector regression model for modeling, permeability prediction of hydrocarbon reservoir. J. Pet. Sci. Eng. 2017 , 150 , 43–53. [ Google Scholar ] [ CrossRef ]
- Chandler, J.; White, D.; Techera, E.J.; Gourvenec, S.; Draper, S. Engineering and legal considerations for decommissioning of offshore oil and gas infrastructure in Australia. Ocean Eng. 2017 , 131 , 338–347. [ Google Scholar ] [ CrossRef ]
- Tian, X.; Liu, Y.; Lin, R.; Sun, P.; Ji, R. An autonomous robot for casing cutting in oil platform decommission. Int. J. Control Autom. 2013 , 6 , 9–20. [ Google Scholar ] [ CrossRef ]
- Wheeler, C.; Hewitt, G. UK offshore decommissioning current Problems. Int. Energy Law Tax. Rev. 2000 , 252 , 18. [ Google Scholar ]
- Paterson, J.; Gordon, G.; Usenmez, E. Oil and Gas Law: Current Practice and Emerging Trends ; Dundee University Press: Dundee, UK, 2011; p. 286. [ Google Scholar ]
- Hekelaar, S.; Gibson, K.; Desai, P. Increasing reliability of cutting/pulling casing in a single trip. SPE Offshore Eur. Oil Gas Conf. Exhib. 2011 , 1 , 367–377. [ Google Scholar ]
- Kaiser, M.J.; Liu, M.M. Decommissioning cost estimation for deepwater floating structures in the US Gulf of Mexico. Ships Offshore Struct. 2015 , 10 , 436–455. [ Google Scholar ] [ CrossRef ]
- Wang, R.H.; Li, L.P.; Zhou, W.D.; Li, H.Z. Experiment and mathematical model of rotary cutting of casing with abrasive water jet. Zhongguo Shiyou Daxue Xuebao 2010 , 34 , 56–59. [ Google Scholar ]
- Yang, H.; Sun, Q.L.; Feng, D.; Zhou, L.; Wang, Z.; Chen, Z. Design of hydraulic cutting device for deepwater casing cutting. Sci. Technol. Eng. 2018 , 18 , 45–50. [ Google Scholar ]
- Dong, H.J.; Lv, Z.G.; Gong, W.M.; Lin, L. Development and application of Model ND—S114 casing internal cutter. China Pet. Mach. 1999 , 27 , 34–35. [ Google Scholar ]
- Liu, H.M.; Ye, H.; Tian, M.; Wu, G.Z.; Xiao, B.J. Application of the mechanical cutting technology for coiled tubing in Yong 25-11 well. Complex Hydrocarb. Reserv. 2014 , 7 , 79–81. [ Google Scholar ]
- Jin, G.; Liu, X.H.; Wu, H.; Cheng, L.; He, L.; Zhang, T. Research on cutting failure of mechanical internal cutter based on ADAMS. J. Saf. Sci. Technol. 2017 , 13 , 99–103. [ Google Scholar ]
- Mamedov, A.A.; Samedov, Y.F. Determination of regime parameters for the work of a pipe cutter when pipes are cut from multistage casing strings. Khimicheskoe I Neftegazov. Mashinostroenie 1994 , 2 , 18–20. [ Google Scholar ]
- Zhou, W.D.; Wang, R.H.; Yang, Y.Y. Models for calculating cutting depth of casing by premixing abrasive water jet. Shiyou Daxue Xuebao J. Univ. Pet. China 2001 , 25 , 3–5. [ Google Scholar ]
- Toulouse, J.E. Improved window cutting efficiency cuts casing exits costs. Thriving on Volatility. In Proceedings of the SPE Asia Pacific Oil and Gas Conference and Exhibition, Adelaide, Australia, 11–13 September 2006; Volume 2, pp. 1046–1049. [ Google Scholar ]
- Ivanov, A.N.; Karapetov, R.V. Selection of the methods to cut the subsea casing assembly during offshore wells abandonment. Ocean Eng. Equip. Technol. 2013 , 173 , 478–491. [ Google Scholar ]
- Zhou, X.; He, S.M.; Tang, M. Structural design and mathematical cutting model of hydraulic section milling for high-strength casings in small wellbores. J. Pet. Sci. Eng. 2022 , 218 , 111044. [ Google Scholar ] [ CrossRef ]
- Chen, Y.; Qin, H.Z.; Tan, S. Resign and research on cutting and retrieving tool of deepwater subsea wellhead system. China Pet. Mach. 2016 , 44 , 70–75. [ Google Scholar ]
- Feng, D.; Huang, K.; Wang, P.; Zhang, H.; Shi, L. Analysis of the eccentric scenario in casing-cutting operations for offshore well abandonment. China Offshore Oil Gas 2017 , 29 , 103–108. [ Google Scholar ]
- YAN, B.; Xia, C.Y.; Wang, P.; Lu, H.B.; Huang, K. Analysis of influencing factors of torque in casing cutting in deep water. Oil Field Equip. 2015 , 44 , 4–7. [ Google Scholar ]
- Feng, D.; Sun, Q.L.; Xia, C.Y.; Wang, J. A Hydraulic Internal Cutter for Deep Sea Casing Cutting. Chinese Patent CN201510116649.4[P], 17 June 2015. [ Google Scholar ]
- He, J.M. Mill Calculation and Common Table ; Mechanical Industry Press: Beijing, China, 2004; pp. 73–99. [ Google Scholar ]
- Charges, R.; Charges, F.; Charges, L.S. An Assessment of Techniques for Removing Offshore Structures ; National Academy Press: Washington, DC, USA, 1996; pp. 47–68. [ Google Scholar ]
- Zhang, Y.Z.; Huang, X.H. Simulation and research of aerospace material milling based on ABAQUS. J. Phys. Conf. Ser. IOP Publ. 2020 , 1653 , 012070. [ Google Scholar ] [ CrossRef ]
- Montassar, F.; Frédéric, M.; Johanna, S.; Rubio, W. Cutting parameters and tool geometry selection for plunge milling - Analysis of cutting forces at the bottom of deep titanium workpieces. J. Manuf. Process. 2021 , 62 , 491–500. [ Google Scholar ] [ CrossRef ]
- Zheng, F.; Han, X.; Lin, H.; Zhao, W. Research on the cutting dynamics for face-milling of spiral bevel gears. Mech. Syst. Signal Process. 2021 , 153 , 107488. [ Google Scholar ] [ CrossRef ]
- Ning, H.F.; Zheng, H.L.; Wang, G.X. Establishment of Analytical Model for CFRP Cutting Force Considering the Radius of the Edge Circle. Materials 2022 , 15 , 2127. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Sun, Q.L.; Xia, L.; Deng, L.; Wang, J.G.; Wang, G.L.; Feng, D. Experimental and Numerical Simulation Analyses of Elbow Erosion in Surface Process of Deepwater Gas Well Testing. J. Fail. Anal. Prev. 2024 , 24 , 202–215. [ Google Scholar ] [ CrossRef ]
- Tang, M.; Li, H.; He, S.; Miao, J.; Huang, B.; He, L.; Zhou, X. Research and Development of High-Performance Section Milling Tool for Φ139.7-mm Casing. Arab. J. Sci. Eng. 2022 , 47 , 12205–12222. [ Google Scholar ] [ CrossRef ]
- Bondor, P.; Rusu, D.O.; Voina, I.D.; Contiu, G.; Popa, M.S. Influence of Variation of Cutting Speed on Wear, Cutting Forces and Tool Temperature during Performance Drilling. Teh. Vjesn. 2023 , 30 , 354–360. [ Google Scholar ]
- Bosire, R.N.; Muvengei, O.M.; Mutua, J.M.; Kimotho, J.K. Finite element based model for predicting induced residual stresses and cutting forces in AISI 1020 steel alloy. Mater. Werkst. 2023 , 54 , 554–570. [ Google Scholar ] [ CrossRef ]
- Zhu, X.H.; Zhou, B.W.; Jing, J.; Shi, J.; Qin, R.Y. Modeling and Optimization of Mechanical Cutting of Downhole Tubing. SPE J. 2024 , 29 , 798–814. [ Google Scholar ] [ CrossRef ]
Click here to enlarge figure
| A/MPa | B/MPa | C | n | m |
|---|---|---|---|---|
| 1150 | 739 | 0.014 | 0.26 | 1.03 |
| Parameters | Value | Parameters | Value |
|---|---|---|---|
| /mm | 151 | f | 3.5 |
| /mm | 168 | 32 | |
| qm/(Kg/m) | 122 | n/(r/min) | 45 |
| K | 3 | /( ) | 85 |
| ρ/(kg/m ) | 1025 | L/m | 850 |
| g/(N/Kg) | 9.8 | 313 | |
| Sz/mm | 0.12 | 340 | |
| Z | 12 | 298 |
| The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
Share and Cite
Sun, Q.; Tian, J.; Jin, Y.; Feng, D.; Hou, L. Modeling and Research on Offshore Casing Cutting of Hydraulic Internal Cutting Device. J. Mar. Sci. Eng. 2024 , 12 , 1026. https://doi.org/10.3390/jmse12061026
Sun Q, Tian J, Jin Y, Feng D, Hou L. Modeling and Research on Offshore Casing Cutting of Hydraulic Internal Cutting Device. Journal of Marine Science and Engineering . 2024; 12(6):1026. https://doi.org/10.3390/jmse12061026
Sun, Qiaolei, Jie Tian, Yujie Jin, Ding Feng, and Lingxia Hou. 2024. "Modeling and Research on Offshore Casing Cutting of Hydraulic Internal Cutting Device" Journal of Marine Science and Engineering 12, no. 6: 1026. https://doi.org/10.3390/jmse12061026
Article Metrics
Article access statistics, further information, mdpi initiatives, follow mdpi.

Subscribe to receive issue release notifications and newsletters from MDPI journals

IMAGES
VIDEO
COMMENTS
Common types of limitations and their ramifications include: Theoretical: limits the scope, depth, or applicability of a study. Methodological: limits the quality, quantity, or diversity of the data. Empirical: limits the representativeness, validity, or reliability of the data. Analytical: limits the accuracy, completeness, or significance of ...
In research, studies can have limitations such as limited scope, researcher subjectivity, and lack of available research tools. Acknowledging the limitations of your study should be seen as a strength. It demonstrates your willingness for transparency, humility, and submission to the scientific method and can bolster the integrity of the study.
Limitation #3: Sample Size & Composition. As we've discussed before, the size and representativeness of your sample are crucial, especially in quantitative research where the robustness of your conclusions often depends on these factors.All too often though, students run into issues achieving a sufficient sample size and composition. To ensure adequacy in terms of your sample size, it's ...
Some Examples of Limitations in Research are as follows: Example 1: Research Title: ... Generally, limitations should be discussed in the conclusion section of a research paper or thesis, although they may also be mentioned in other sections, such as the introduction or methods. The specific limitations that are discussed will depend on the ...
how the study enables future research—will help ensure that the study's drawbacks are not the last thing reviewers read in the paper. Start this "limitations" paragraph with a simple topic sentence that signals what you're about to discuss. For example: "Our study had some limitations."
Sample Size Limitations in Qualitative Research. Sample sizes are typically smaller in qualitative research because, as the study goes on, acquiring more data does not necessarily lead to more information. This is because one occurrence of a piece of data, or a code, is all that is necessary to ensure that it becomes part of the analysis framework.
Step 1. Identify the limitation (s) of the study. This part should comprise around 10%-20% of your discussion of study limitations. The first step is to identify the particular limitation (s) that affected your study. There are many possible limitations of research that can affect your study, but you don't need to write a long review of all ...
Here's an example of a limitation explained in a research paper about the different options and emerging solutions for delaying memory decline. These statements appeared in the first two sentences of the discussion section: "Approaches like stem cell transplantation and vaccination in AD [Alzheimer's disease] work on a cellular or molecular level in the laboratory.
The ideal way is to divide your limitations section into three steps: 1. Identify the research constraints; 2. Describe in great detail how they affect your research; 3. Mention the opportunity for future investigations and give possibilities. By following this method while addressing the constraints of your research, you will be able to ...
3. Identify your limitations of research and explain their importance. 4. Provide the necessary depth, explain their nature, and justify your study choices. 5. Write how you are suggesting that it is possible to overcome them in the future. Limitations can help structure the research study better.
Limitations of a dissertation are potential weaknesses in your study that are mostly out of your control, given limited funding, choice of research design, statistical model constraints, or other factors. In addition, a limitation is a restriction on your study that cannot be reasonably dismissed and can affect your design and results.
For example, in their 2021 Cell Reports study on macrophage polarization mechanisms, dermatologist Alexander Marneros and colleagues wrote the following. 1. A limitation of studying macrophage polarization in vitro is that this approach only partially captures the tissue microenvironment context in which many different factors affect macrophage polarization.
2.3. Limitations Example 3. It is important to remember not to end your paper with limitations. Finish your paper on a positive note by telling your readers about the benefits of your research and possible future directions. In the following example, right after listing the limitations, the authors proceed to talk about the positive aspects of ...
Information about the limitations of your study are generally placed either at the beginning of the discussion section of your paper so the reader knows and understands the limitations before reading the rest of your analysis of the findings, or, the limitations are outlined at the conclusion of the discussion section as an acknowledgement of the need for further study.
Limitations are usually listed at the end of your Discussion section, though they can also be added throughout. Especially for a long manuscript or for an essay or dissertation, the latter may be useful for the reader. Writing on your limitations: Words and structure. This study did have some limitations. Three notable limitations affected this ...
Possible Methodological Limitations. Sample size-- the number of the units of analysis you use in your study is dictated by the type of research problem you are investigating. Note that, if your sample size is too small, it will be difficult to find significant relationships from the data, as statistical tests normally require a larger sample ...
Research Limitations. Research limitations are, at the simplest level, the weaknesses of the study, based on factors that are often outside of your control as the researcher. These factors could include things like time, access to funding, equipment, data or participants.For example, if you weren't able to access a random sample of participants for your study and had to adopt a convenience ...
Well, that depends entirely on the nature of your study. You'll need to comb through your research approach, methodology, testing processes, and expected results to identify the type of limitations your study may be exposed to. It's worth noting that this understanding can only offer a broad idea of the possible restrictions you'll face ...
For example, if conducting a meta-analysis of the secondary data has not been stated as your research objective, no need to mention it as your research limitation. Research limitations in a typical dissertation may relate to the following points: 1. Formulation of research aims and objectives. You might have formulated research aims and ...
There is no "one best way" to structure the Research Limitations section of your dissertation. However, we recommend a structure based on three moves: the announcing, reflecting and forward looking move. The announcing move immediately allows you to identify the limitations of your dissertation and explain how important each of these ...
Writing the limitations of the research papers is often assumed to require lots of effort. However, identifying the limitations of the study can help structure the research better. Therefore, do not underestimate the importance of research study limitations. 3. Opportunity to make suggestions for further research.
conference, or a published research paper in an academic journal. "Limitations of Research". is a section in the standard research report (the research report is usually divided into the ...
Any limitation influences a research paper. It is unknown how much and to what extent any limitation affects other limitations, but it does create a cascading domino effect of ever-increasing interactions that compromise findings and conclusions. ... This is a sample of limitations and a few of their component variables under the rubric of a ...
Begin with a clear statement of the principal findings. This will reinforce the main take-away for the reader and set up the rest of the discussion. Explain why the outcomes of your study are important to the reader. Discuss the implications of your findings realistically based on previous literature, highlighting both the strengths and ...
The term 'lived experience' has its origins in phenomenology, though historically it has been focussed on participants as the 'subject' of research rather than as active contributors throughout the research process (Frechette et al., 2020).Lived experience researchers or co-researchers are defined, for the purposes of this paper, as people who carry out research with knowledge and ...
The sample covers research published in 56 journals. ... Most of these papers suggest hybrid mentoring is positively related to the full range of career development outcomes, ... as well as general sample size limitations, prevented this analysis. Nonetheless, where possible (e.g. mentoring for targeted groups) the review does provide nuanced ...
Antimicrobial resistance is a global threat to public health. Without proactive intervention, common infections may become untreatable, restricting the types of clinical intervention that can be ...
Please note several limitations on length: (1) your abstract should be no more than 150 words, as the abstract will also be used for the conference program, (2) your completed research paper should be no more than 10 pages (approx. 5,000 words, including figures, tables, references, and appendices).
At the beginning of my career, I worked for the health-care-software specialist HBOC. One day, a woman from human resources came into the cafeteria with a roll of tape and began sticking posters ...
The field example shows the limitation of judging casing cutting by a sudden drop in torque. ... Feature papers represent the most advanced research with significant potential for high impact in the field. A Feature Paper should be a substantial original Article that involves several techniques or approaches, provides an outlook for future ...